 3D Periodic Editor 3D Molecular And Reaction Editor Expert Editor Microsoft Quantum Editor EMSL Aerosol Workshop Editor Manual
3D Periodic Editor 3D Molecular And Reaction Editor Expert Editor Microsoft Quantum Editor EMSL Aerosol Workshop Editor Manual Results from an EMSL Arrows Calculation
| EMSL Arrows is a revolutionary approach to materials and chemical simulations that uses NWChem and chemical computational databases to make materials and chemical modeling accessible via a broad spectrum of digital communications including posts to web APIs, social networks, and traditional email. |
Molecular modeling software has previously been extremely complex, making it prohibative to all but experts in the field, yet even experts can struggle to perform calculations. This service is designed to be used by experts and non-experts alike. Experts can carry out and keep track of large numbers of complex calculations with diverse levels of theories present in their workflows. Additionally, due to a streamlined and easy-to-use input, non-experts can carry out a wide variety of molecular modeling calculations previously not accessible to them.
The id(s) for emsiles = O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C theory{dft} xc{b3lyp} basis{6-311++G(2d,2p)} solvation_type{COSMO-SMD:o-cresol} ^{-1} are: 73405
Use id=% instead of esmiles to print other entries.
mformula = C7H4N3O6
iupac = 2-methyl-1,3,5-trinitrobenzene anion
PubChem = 8376
PubChem LCSS = 8376
cas = 118-96-7
synonyms = 2,4,6-TRINITROTOLUENE; Trinitrotoluene; Trotyl; 2-Methyl-1,3,5-trinitrobenzene; 118-96-7; s-Trinitrotoluol; s-Trinitrotoluene; Tolite; Tritol; sym-Trinitrotoluol; trinitrotoluol; Trojnitrotoluen; Gradetol; Tolit; Tnt-tolite; Trotyl oil; alpha-TNT; 2,4,6-Trinitrotoluol; TNT; Tritol (explosive); Benzene, 2-methyl-1,3,5-trinitro-; Toluene, 2,4,6-trinitro-; sym-Trinitrotoluene; NCI-C56155; Trinitrotoluen; 2,4,6-Trinitrotolueen; 1-Methyl-2,4,6-trinitrobenzene; UNII-H43RF5TRM5; NSC 36949; .alpha.-TNT; 2,4,6-TNT; H43RF5TRM5; 2,4,6-trinitrotoluene (tnt); trilit; DTXSID7024372; CHEBI:46053; 2,4,6-trinitritoluene; TNT-tolite [French]; Trojnitrotoluen [Polish]; TNL; CCRIS 1299; HSDB 1146; 2,4,6-Trinitrotolueen [Dutch]; 2,4,6-Trinitrotoluol [German]; EINECS 204-289-6; UN0209; UN1356; alpha-trinitrotoluol; Trinitrotoluene, dry; Trinitrotoluene, wet; 2,6-Trinitrotoluol; 2,6-Trinitrotolueen; 2,6-Trinitrotoluene; 2,4,6-trinitro-toluene; EC 204-289-6; SCHEMBL20676; UN 0209 (Salt/Mix); UN 1356 (Salt/Mix); WLN: WNR B1 CNW ENW; 2-Methyl-1,5-trinitrobenzene; Toluene,4,6-trinitro- (wet); CHEMBL1236345; SCHEMBL12305492; 1-methyl-2,4,6-trinitrotoluene; NSC36949; Trinitrotoluene or TNT, dry or wetted with < 30% water, by mass; NSC-36949; ZINC14880028; AKOS001092689; DB01676; MCULE-8164226079; Q170167; Trinitrotoluene, wetted with not <30% water, by mass; Z57056414; 2,4,6-Trinitrotoluene (TNT) 10 microg/mL in Cyclohexane; 2,4,6-Trinitrotoluene (TNT) 100 microg/mL in Cyclohexane; Trinitrotoluene or TNT, dry or wetted with <30% water, by mass; Trinitrotoluene, wetted with not <30% water, by mass [UN1356] [Flammable solid]; 2,4,6-Trinitrotoluene solution, 10 mg/mL in acetonitrile, ampule of 5 mL, certified reference material; 2,4,6-Trinitrotoluene solution, 1000 mug/mL in acetonitrile, ampule of 1.2 mL, certified reference material; Trinitrotoluene or TNT, dry or wetted with <30% water, by mass [UN0209] [Explosive 1.1D]
Search Links to Other Online Resources (may not be available):
- Google Structure Search
- EPA CompTox Database
- Chemical Entities of Biological Interest (ChEBI)
- NIH ChemIDplus - A TOXNET DATABASE
- The Human Metabolome Database (HMDB)
- OECD eChemPortal
- Google Scholar
+==================================================+
|| Molecular Calculation ||
+==================================================+
Id = 73405
NWOutput = Link to NWChem Output (download)
Datafiles:
homo-restricted.cube-21231-2022-5-7-16:1:28 (download)
lumo-restricted.cube-21231-2022-5-7-16:1:28 (download)
mo_orbital_nwchemarrows.out-156812-2022-8-3-8:41:41 (download)
image_resset: api/image_reset/73405
Calculation performed by Eric Bylaska - constance.pnl.gov
Numbers of cpus used for calculation = 48
Calculation walltime = 17326.800000 seconds (0 days 4 hours 48 minutes 46 seconds)
+----------------+
| Energetic Data |
+----------------+
Id = 73405
iupac = 2-methyl-1,3,5-trinitrobenzene anion
mformula = C7H4N3O6
inchi = InChI=1S/C7H4N3O6/c1-4-6(9(13)14)2-5(8(11)12)3-7(4)10(15)16/h2H,1H3
inchikey = DWRNYJRBGMEGOX-UHFFFAOYSA-N
esmiles = O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C theory{dft} xc{b3lyp} basis{6-311++G(2d,2p)} solvation_type{COSMO-SMD:o-cresol} ^{-1}
calculation_type = ovc
theory = dft
xc = b3lyp
basis = 6-311++G(2d,2p)
charge,mult = -1 1
energy = -884.776698 Hartrees
enthalpy correct.= 0.134236 Hartrees
entropy = 115.005 cal/mol-K
solvation energy = -34.040 kcal/mol solvation_type = COSMO-SMD:o-cresol
Sitkoff cavity dispersion = 2.756 kcal/mol
Honig cavity dispersion = 9.481 kcal/mol
ASA solvent accesible surface area = 379.225 Angstrom2
ASA solvent accesible volume = 366.299 Angstrom3
+-----------------+
| Structural Data |
+-----------------+
JSmol: an open-source HTML5 viewer for chemical structures in 3D
Number of Atoms = 20
Units are Angstrom for bonds and degrees for angles
Type I J K L M Value
----------- ----- ----- ----- ----- ----- ----------
1 Stretch O1 N2 1.23656
2 Stretch N2 O3 1.21849
3 Stretch N2 C4 1.50731
4 Stretch C4 C5 1.38731
5 Stretch C4 C9 1.38514
6 Stretch C5 C6 1.38511
7 Stretch C6 C7 1.39411
8 Stretch C6 N14 1.51130
9 Stretch C7 C8 1.41399
10 Stretch C7 C13 1.50685
11 Stretch C8 C9 1.39090
12 Stretch C8 N10 1.45834
13 Stretch C9 H17 1.07682
14 Stretch N10 O11 1.23336
15 Stretch N10 O12 1.23172
16 Stretch C13 H18 1.08657
17 Stretch C13 H19 1.08954
18 Stretch C13 H20 1.08767
19 Stretch N14 O15 1.22520
20 Stretch N14 O16 1.22355
21 Bend O1 N2 O3 123.48669
22 Bend O1 N2 C4 117.59754
23 Bend O3 N2 C4 118.91469
24 Bend N2 C4 C5 118.96199
25 Bend N2 C4 C9 114.98862
26 Bend C5 C4 C9 126.04938
27 Bend C4 C5 C6 110.83207
28 Bend C5 C6 C7 130.03003
29 Bend C5 C6 N14 114.70810
30 Bend C7 C6 N14 115.26094
31 Bend C6 C7 C8 113.35971
32 Bend C6 C7 C13 122.30178
33 Bend C8 C7 C13 124.30781
34 Bend C7 C8 C9 121.76397
35 Bend C7 C8 N10 121.88106
36 Bend C9 C8 N10 116.35260
37 Bend C4 C9 C8 117.95108
38 Bend C4 C9 H17 122.26657
39 Bend C8 C9 H17 119.77746
40 Bend C8 N10 O11 118.75682
41 Bend C8 N10 O12 118.29199
42 Bend O11 N10 O12 122.94577
43 Bend C7 C13 H18 110.79041
44 Bend C7 C13 H19 110.88124
45 Bend C7 C13 H20 111.39058
46 Bend H18 C13 H19 108.47251
47 Bend H18 C13 H20 108.53642
48 Bend H19 C13 H20 106.62229
49 Bend C6 N14 O15 117.62039
50 Bend C6 N14 O16 117.97502
51 Bend O15 N14 O16 124.40369
52 Dihedral O1 N2 C4 C5 169.03124
53 Dihedral O1 N2 C4 C9 -10.94008
54 Dihedral N2 C4 C5 C6 -179.46407
55 Dihedral N2 C4 C9 C8 178.72270
56 Dihedral N2 C4 C9 H17 -0.47026
57 Dihedral O3 N2 C4 C5 -11.33357
58 Dihedral O3 N2 C4 C9 168.69511
59 Dihedral C4 C5 C6 C7 0.66164
60 Dihedral C4 C5 C6 N14 -179.71026
61 Dihedral C4 C9 C8 C7 0.89925
62 Dihedral C4 C9 C8 N10 -178.55060
63 Dihedral C5 C4 C9 C8 -1.24626
64 Dihedral C5 C4 C9 H17 179.56078
65 Dihedral C5 C6 C7 C8 -0.92874
66 Dihedral C5 C6 C7 C13 -178.99362
67 Dihedral C5 C6 N14 O15 -95.98747
68 Dihedral C5 C6 N14 O16 83.68161
69 Dihedral C6 C5 C4 C9 0.50377
70 Dihedral C6 C7 C8 C9 0.06401
71 Dihedral C6 C7 C8 N10 179.48345
72 Dihedral C6 C7 C13 H18 15.76403
73 Dihedral C6 C7 C13 H19 -104.73895
74 Dihedral C6 C7 C13 H20 136.68977
75 Dihedral C7 C6 N14 O15 83.69765
76 Dihedral C7 C6 N14 O16 -96.63326
77 Dihedral C7 C8 C9 H17 -179.88698
78 Dihedral C7 C8 N10 O11 -22.93456
79 Dihedral C7 C8 N10 O12 157.88739
80 Dihedral C8 C7 C6 N14 179.44483
81 Dihedral C8 C7 C13 H18 -162.08519
82 Dihedral C8 C7 C13 H19 77.41182
83 Dihedral C8 C7 C13 H20 -41.15946
84 Dihedral C9 C8 C7 C13 178.08384
85 Dihedral C9 C8 N10 O11 156.51458
86 Dihedral C9 C8 N10 O12 -22.66346
87 Dihedral N10 C8 C7 C13 -2.49672
88 Dihedral N10 C8 C9 H17 0.66317
89 Dihedral C13 C7 C6 N14 1.37995
+-----------------+
| Eigenvalue Data |
+-----------------+
Id = 73405
iupac = 2-methyl-1,3,5-trinitrobenzene anion
mformula = C7H4N3O6
InChI = InChI=1S/C7H4N3O6/c1-4-6(9(13)14)2-5(8(11)12)3-7(4)10(15)16/h2H,1H3
smiles = O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C
esmiles = O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C theory{dft} xc{b3lyp} basis{6-311++G(2d,2p)} solvation_type{COSMO-SMD:o-cresol} ^{-1}
theory = dft
xc = b3lyp
basis = 6-311++G(2d,2p)
charge = -1
mult = 1
solvation_type = COSMO-SMD:o-cresol
twirl webpage = TwirlMol Link
image webpage = GIF Image Link
Eigevalue Spectra
---- ---- 67.92 eV
---- ----
-- -- -- -
---- ----
-- -- -- -
--- -- ---
---- ----
6 - - - -
-- -- -- -
--- -- ---
7 - - - -
- - - - --
10 - - - -
- - - - --
8 - - - -
10 - - - -
10 - - - -
13 - - - -
9 - - - -
6 - - - -
6 - - - -
7 - - - -
8 - - - -
9 - - - -
7 - - - -
11 - - - -
9 - - - -
14 - - - -
15 - - - -
16 - - - -
16 - - - -
9 - - - -
--- -- ---
--- -- --- LUMO= -2.74 eV
HOMO= -5.75 eV +++ ++ +++
9 + + + +
++ ++ ++ +
++ ++ ++ +
8 + + + +
++++ ++++
++++ ++++
++++++++++
++++ ++++
++++++++++
+++ ++ +++
-33.76 eV +++ ++ +++
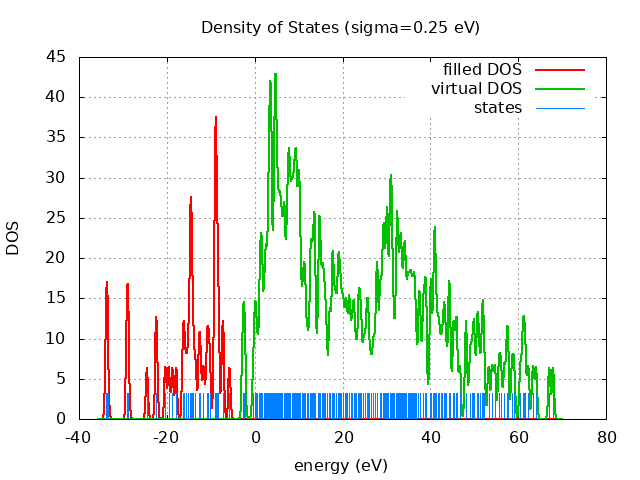
spin eig occ ---------------------------- restricted -33.76 2.00 restricted -33.73 2.00 restricted -33.49 2.00 restricted -29.08 2.00 restricted -28.97 2.00 restricted -28.77 2.00 restricted -24.55 2.00 restricted -22.43 2.00 restricted -22.38 2.00 restricted -20.35 2.00 restricted -19.57 2.00 restricted -18.75 2.00 restricted -17.89 2.00 restricted -16.31 2.00 restricted -16.10 2.00 restricted -15.62 2.00 restricted -15.11 2.00 restricted -14.80 2.00 restricted -14.55 2.00 restricted -14.51 2.00 restricted -14.47 2.00 restricted -14.30 2.00 restricted -13.94 2.00 restricted -13.47 2.00 restricted -12.66 2.00 restricted -12.37 2.00 restricted -11.66 2.00 restricted -10.92 2.00 restricted -10.61 2.00 restricted -10.25 2.00 restricted -9.13 2.00 restricted -9.04 2.00 restricted -8.94 2.00 restricted -8.91 2.00 restricted -8.80 2.00 restricted -8.78 2.00 restricted -8.57 2.00 restricted -8.37 2.00 restricted -8.24 2.00 restricted -7.32 2.00 restricted -7.18 2.00 restricted -5.75 2.00 restricted -2.74 0.00 restricted -2.47 0.00 restricted -2.27 0.00 restricted -0.29 0.00 restricted 0.11 0.00 restricted 0.24 0.00 restricted 0.64 0.00 restricted 1.07 0.00 restricted 1.26 0.00 restricted 1.43 0.00 restricted 1.59 0.00 restricted 1.69 0.00 restricted 2.04 0.00 restricted 2.28 0.00 restricted 2.54 0.00 restricted 2.58 0.00 restricted 2.81 0.00 restricted 3.10 0.00 restricted 3.19 0.00 restricted 3.30 0.00 restricted 3.45 0.00 restricted 3.49 0.00 restricted 3.56 0.00 restricted 3.70 0.00 restricted 3.70 0.00 restricted 3.84 0.00 restricted 3.92 0.00 restricted 4.27 0.00 restricted 4.46 0.00 restricted 4.55 0.00 restricted 4.62 0.00 restricted 4.63 0.00 restricted 4.73 0.00 restricted 4.86 0.00 restricted 4.95 0.00 restricted 5.00 0.00 restricted 5.18 0.00 restricted 5.31 0.00 restricted 5.48 0.00 restricted 5.62 0.00 restricted 5.74 0.00 restricted 5.90 0.00 restricted 6.02 0.00 restricted 6.21 0.00 restricted 6.29 0.00 restricted 6.57 0.00 restricted 6.70 0.00 restricted 6.74 0.00 restricted 6.89 0.00 restricted 7.22 0.00 restricted 7.29 0.00 restricted 7.55 0.00 restricted 7.58 0.00 restricted 7.72 0.00 restricted 7.80 0.00 restricted 7.97 0.00 restricted 8.05 0.00 restricted 8.14 0.00 restricted 8.44 0.00 restricted 8.49 0.00 restricted 8.51 0.00 restricted 8.78 0.00 restricted 8.89 0.00 restricted 8.99 0.00 restricted 9.13 0.00 restricted 9.30 0.00 restricted 9.36 0.00 restricted 9.45 0.00 restricted 9.58 0.00 restricted 9.74 0.00 restricted 9.91 0.00 restricted 10.11 0.00 restricted 10.15 0.00 restricted 10.20 0.00 restricted 10.35 0.00 restricted 10.68 0.00 restricted 10.90 0.00 restricted 11.16 0.00 restricted 11.34 0.00 restricted 11.48 0.00 restricted 11.82 0.00 restricted 12.16 0.00 restricted 12.61 0.00 restricted 12.71 0.00 restricted 12.90 0.00 restricted 13.06 0.00 restricted 13.27 0.00 restricted 13.50 0.00 restricted 13.59 0.00 restricted 13.69 0.00 restricted 13.84 0.00 restricted 14.39 0.00 restricted 14.62 0.00 restricted 14.74 0.00 restricted 14.83 0.00 restricted 14.98 0.00 restricted 15.29 0.00 restricted 15.46 0.00 restricted 15.65 0.00 restricted 15.89 0.00 restricted 16.10 0.00 restricted 16.42 0.00 restricted 17.03 0.00 restricted 17.12 0.00 restricted 17.63 0.00 restricted 17.74 0.00 restricted 17.83 0.00 restricted 18.12 0.00 restricted 18.42 0.00 restricted 18.58 0.00 restricted 18.97 0.00 restricted 19.10 0.00 restricted 19.19 0.00 restricted 19.54 0.00 restricted 19.65 0.00 restricted 20.01 0.00 restricted 20.19 0.00 restricted 20.52 0.00 restricted 20.82 0.00 restricted 21.00 0.00 restricted 21.49 0.00 restricted 21.51 0.00 restricted 21.92 0.00 restricted 22.26 0.00 restricted 22.54 0.00 restricted 22.75 0.00 restricted 23.17 0.00 restricted 23.61 0.00 restricted 23.75 0.00 restricted 24.05 0.00 restricted 24.28 0.00 restricted 24.75 0.00 restricted 25.11 0.00 restricted 25.50 0.00 restricted 25.73 0.00 restricted 25.96 0.00 restricted 26.45 0.00 restricted 26.96 0.00 restricted 27.31 0.00 restricted 27.77 0.00 restricted 27.90 0.00 restricted 27.92 0.00 restricted 28.43 0.00 restricted 28.62 0.00 restricted 28.81 0.00 restricted 29.19 0.00 restricted 29.24 0.00 restricted 29.36 0.00 restricted 29.55 0.00 restricted 29.79 0.00 restricted 29.99 0.00 restricted 30.18 0.00 restricted 30.20 0.00 restricted 30.31 0.00 restricted 30.77 0.00 restricted 30.83 0.00 restricted 30.91 0.00 restricted 31.04 0.00 restricted 31.20 0.00 restricted 31.30 0.00 restricted 31.50 0.00 restricted 31.84 0.00 restricted 32.28 0.00 restricted 32.33 0.00 restricted 32.47 0.00 restricted 32.55 0.00 restricted 32.83 0.00 restricted 33.03 0.00 restricted 33.22 0.00 restricted 33.35 0.00 restricted 33.49 0.00 restricted 33.79 0.00 restricted 33.99 0.00 restricted 34.19 0.00 restricted 34.27 0.00 restricted 34.51 0.00 restricted 34.83 0.00 restricted 34.94 0.00 restricted 35.23 0.00 restricted 35.42 0.00 restricted 35.63 0.00 restricted 35.87 0.00 restricted 36.09 0.00 restricted 36.35 0.00 restricted 36.48 0.00 restricted 36.78 0.00 restricted 37.29 0.00 restricted 37.56 0.00 restricted 37.67 0.00 restricted 38.27 0.00 restricted 38.39 0.00 restricted 38.76 0.00 restricted 38.91 0.00 restricted 39.10 0.00 restricted 39.91 0.00 restricted 40.12 0.00 restricted 40.21 0.00 restricted 40.71 0.00 restricted 40.90 0.00 restricted 40.99 0.00 restricted 41.11 0.00 restricted 41.34 0.00 restricted 41.69 0.00 restricted 41.97 0.00 restricted 42.35 0.00 restricted 42.74 0.00 restricted 43.08 0.00 restricted 43.27 0.00 restricted 43.64 0.00 restricted 44.14 0.00 restricted 44.39 0.00 restricted 44.42 0.00 restricted 45.18 0.00 restricted 45.41 0.00 restricted 45.88 0.00 restricted 46.05 0.00 restricted 46.72 0.00 restricted 48.04 0.00 restricted 48.18 0.00 restricted 48.97 0.00 restricted 49.39 0.00 restricted 49.82 0.00 restricted 50.04 0.00 restricted 50.65 0.00 restricted 50.79 0.00 restricted 51.24 0.00 restricted 51.79 0.00 restricted 51.93 0.00 restricted 52.25 0.00 restricted 53.27 0.00 restricted 54.06 0.00 restricted 54.68 0.00 restricted 55.60 0.00 restricted 56.07 0.00 restricted 56.84 0.00 restricted 57.43 0.00 restricted 57.66 0.00 restricted 58.58 0.00 restricted 59.06 0.00 restricted 60.51 0.00 restricted 61.03 0.00 restricted 61.29 0.00 restricted 61.62 0.00 restricted 62.29 0.00 restricted 63.37 0.00 restricted 64.05 0.00 restricted 67.07 0.00 restricted 67.92 0.00
+----------------------------------------+ | Vibrational Density of States Analysis | +----------------------------------------+ Total number of frequencies = 60 Total number of negative frequencies = 1 - w_negative = -69.7 cm-1 Number of lowest frequencies = 16 (frequency threshold = 500 ) Exact dos norm = 54.000 Generating vibrational DOS Generating model vibrational DOS to have a proper norm 10.00 53.01 16.00 54.00 50.00 52.62 15.62 54.00 100.00 51.85 14.85 54.00 Generating IR Spectra Writing vibrational density of states (DOS) to vdos.dat Writing model vibrational density of states (DOS_FIXED) to vdos-model.dat Writing IR spectra to irdos.dat Temperature= 298.15 zero-point correction to energy = 75.547 kcal/mol ( 0.120392) vibrational contribution to enthalpy correction = 81.865 kcal/mol ( 0.130461) vibrational contribution to Entropy = 40.219 cal/mol-k hindered rotor enthalpy correction = 0.000 kcal/mol ( 0.000000) hindered rotor entropy correction = 0.000 cal/mol-k
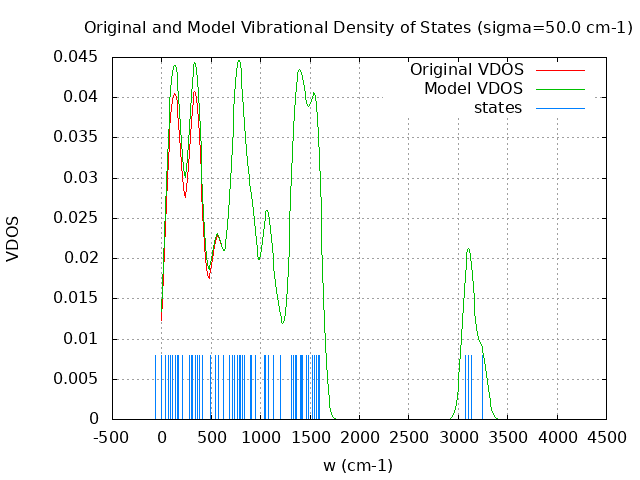
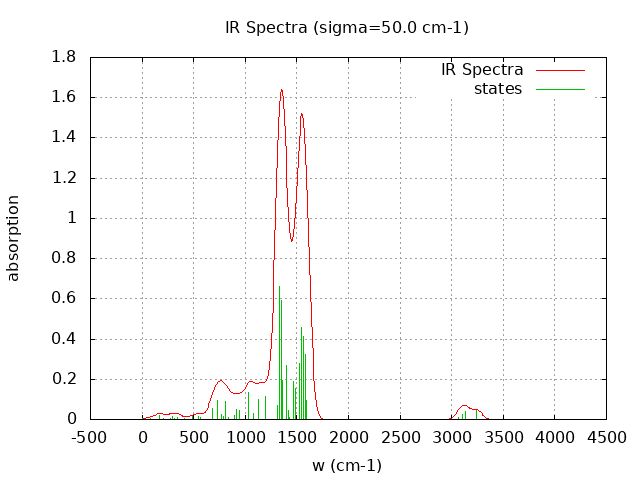
DOS sigma = 10.000000
- vibrational DOS enthalpy correction = 0.130464 kcal/mol ( 81.867 kcal/mol)
- model vibrational DOS enthalpy correction = 0.131533 kcal/mol ( 82.538 kcal/mol)
- vibrational DOS Entropy = 0.000064 ( 40.365 cal/mol-k)
- model vibrational DOS Entropy = 0.000068 ( 42.605 cal/mol-k)
- original gas Energy = -884.776698 (-555205.756 kcal/mol)
- original gas Enthalpy = -884.642462 (-555121.522 kcal/mol, delta= 0.000)
- unajusted DOS gas Enthalpy = -884.642459 (-555121.520 kcal/mol, delta= 0.002)
- model DOS gas Enthalpy = -884.641390 (-555120.849 kcal/mol, delta= 0.673)
- original gas Entropy = 0.000183 ( 115.005 cal/mol-k,delta= 0.000)
- unajusted DOS gas Entropy = 0.000184 ( 115.150 cal/mol-k,delta= 0.145)
- model DOS gas Entropy = 0.000187 ( 117.391 cal/mol-k,delta= 2.386)
- original gas Free Energy = -884.697105 (-555155.811 kcal/mol, delta= 0.000)
- unadjusted DOS gas Free Energy = -884.697171 (-555155.852 kcal/mol, delta= -0.041)
- model DOS gas Free Energy = -884.697167 (-555155.849 kcal/mol, delta= -0.039)
- original sol Free Energy = -884.751351 (-555189.851 kcal/mol)
- unadjusted DOS sol Free Energy = -884.751417 (-555189.892 kcal/mol)
- model DOS sol Free Energy = -884.751413 (-555189.889 kcal/mol)
DOS sigma = 50.000000
- vibrational DOS enthalpy correction = 0.130237 kcal/mol ( 81.725 kcal/mol)
- model vibrational DOS enthalpy correction = 0.131735 kcal/mol ( 82.665 kcal/mol)
- vibrational DOS Entropy = 0.000065 ( 40.951 cal/mol-k)
- model vibrational DOS Entropy = 0.000070 ( 44.183 cal/mol-k)
- original gas Energy = -884.776698 (-555205.756 kcal/mol)
- original gas Enthalpy = -884.642462 (-555121.522 kcal/mol, delta= 0.000)
- unajusted DOS gas Enthalpy = -884.642686 (-555121.662 kcal/mol, delta= -0.140)
- model DOS gas Enthalpy = -884.641188 (-555120.722 kcal/mol, delta= 0.800)
- original gas Entropy = 0.000183 ( 115.005 cal/mol-k,delta= 0.000)
- unajusted DOS gas Entropy = 0.000184 ( 115.737 cal/mol-k,delta= 0.732)
- model DOS gas Entropy = 0.000190 ( 118.969 cal/mol-k,delta= 3.964)
- original gas Free Energy = -884.697105 (-555155.811 kcal/mol, delta= 0.000)
- unadjusted DOS gas Free Energy = -884.697676 (-555156.169 kcal/mol, delta= -0.358)
- model DOS gas Free Energy = -884.697714 (-555156.193 kcal/mol, delta= -0.382)
- original sol Free Energy = -884.751351 (-555189.851 kcal/mol)
- unadjusted DOS sol Free Energy = -884.751922 (-555190.209 kcal/mol)
- model DOS sol Free Energy = -884.751960 (-555190.233 kcal/mol)
DOS sigma = 100.000000
- vibrational DOS enthalpy correction = 0.129776 kcal/mol ( 81.435 kcal/mol)
- model vibrational DOS enthalpy correction = 0.132153 kcal/mol ( 82.927 kcal/mol)
- vibrational DOS Entropy = 0.000061 ( 38.581 cal/mol-k)
- model vibrational DOS Entropy = 0.000069 ( 43.499 cal/mol-k)
- original gas Energy = -884.776698 (-555205.756 kcal/mol)
- original gas Enthalpy = -884.642462 (-555121.522 kcal/mol, delta= 0.000)
- unajusted DOS gas Enthalpy = -884.643147 (-555121.952 kcal/mol, delta= -0.430)
- model DOS gas Enthalpy = -884.640770 (-555120.460 kcal/mol, delta= 1.062)
- original gas Entropy = 0.000183 ( 115.005 cal/mol-k,delta= 0.000)
- unajusted DOS gas Entropy = 0.000181 ( 113.367 cal/mol-k,delta= -1.638)
- model DOS gas Entropy = 0.000188 ( 118.285 cal/mol-k,delta= 3.280)
- original gas Free Energy = -884.697105 (-555155.811 kcal/mol, delta= 0.000)
- unadjusted DOS gas Free Energy = -884.697012 (-555155.752 kcal/mol, delta= 0.059)
- model DOS gas Free Energy = -884.696971 (-555155.726 kcal/mol, delta= 0.084)
- original sol Free Energy = -884.751351 (-555189.851 kcal/mol)
- unadjusted DOS sol Free Energy = -884.751258 (-555189.792 kcal/mol)
- model DOS sol Free Energy = -884.751217 (-555189.766 kcal/mol)
Normal Mode Frequency (cm-1) IR Intensity (arbitrary)
----------- ---------------- ------------------------
1 -69.700 0.088
2 -0.000 0.077
3 -0.000 0.659
4 -0.000 0.312
5 0.000 0.326
6 0.000 0.214
7 0.000 0.905
8 40.440 0.139
9 61.950 0.101
10 83.970 0.672
11 110.940 0.223
12 141.650 0.030
13 156.170 0.264
14 173.050 2.475
15 203.490 0.601
16 276.740 0.347
17 297.280 1.803
18 310.410 0.387
19 341.340 1.553
20 358.900 0.073
21 377.410 0.254
22 409.090 0.329
23 487.990 1.563
24 545.680 1.616
25 567.910 0.968
26 619.830 0.039
27 682.620 6.871
28 719.330 0.185
29 732.610 11.677
30 766.580 3.045
31 785.080 1.680
32 793.290 0.800
33 812.700 10.955
34 838.830 1.501
35 898.900 2.527
36 911.800 6.323
37 944.130 5.666
38 1034.810 17.109
39 1053.330 0.267
40 1078.460 3.442
41 1132.720 12.440
42 1199.900 14.304
43 1308.960 8.537
44 1332.210 83.032
45 1353.090 74.427
46 1359.510 24.021
47 1403.450 33.341
48 1417.400 5.712
49 1427.370 1.292
50 1467.020 23.660
51 1487.760 19.581
52 1524.760 35.055
53 1545.640 57.126
54 1568.360 51.854
55 1587.180 40.390
56 1592.640 11.901
57 3068.610 1.341
58 3105.490 3.139
59 3130.630 4.962
60 3241.950 5.821
No Hindered Rotor Data
+-------------------------------------+
| Reactions Contained in the Database |
+-------------------------------------+
Reactions Containing INCHIKEY = DWRNYJRBGMEGOX-UHFFFAOYSA-N
Reactionid Erxn(gas) Hrxn(gas) Grxn(gas) delta_Solv Grxn(aq) ReactionType Reaction
20602 251.137 248.678 245.579 -14.703 230.876 AB + C --> AC + B "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + [SH-] ^{-1} --> O=N(=O)c1[c]c(N(=O)=O)c(c([c]1)N(=O)=O)C + S ^{-2}"
20249 -5.341 -6.593 -10.495 -5.141 -15.636 AB + C --> AC + B "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + [SH-] ^{-1} --> O=N(=O)c1cc(S)c(c([c]1)N(=O)=O)C ^{-1} + O=[N]=O ^{-1}"
20238 251.138 248.678 245.579 -77.683 167.896 AB + C --> AC + B "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + [SH-] ^{-1} --> O=N(=O)c1[c]c(N(=O)=O)c(c([c]1)N(=O)=O)C + S ^{-2}"
20138 8.136 7.791 6.617 -10.201 -3.585 EA + BCD --> AB + CDE "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + O --> O=N(=O)c1cc(O)c(c([c]1)N(=O)=O)C ^{-1} + ON=O"
20125 -44.065 -44.063 -46.408 15.187 -31.222 AB + C --> AC + B "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + hydroxide ^{-1} --> O=N(=O)c1cc(O)c(c([c]1)N(=O)=O)C ^{-1} + O=[N]=O ^{-1}"
19913 255.025 255.474 252.612 -116.296 136.316 AB + C --> AC + B "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + hydroxide ^{-1} --> O=N(=O)c1[c]c(N(=O)=O)c(c([c]1)N(=O)=O)C + O ^{-2}"
19865 20.248 22.346 33.196 -27.217 5.980 A + B --> AB "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + hydroxide ^{-1} --> O=N(=O)C1=C(C)C(=[CH](C(=[C]1)N(=O)=O)O)N(=O)=O ^{-2}"
19860 250.557 251.165 250.234 -116.977 133.257 AB + C --> AC + B "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + hydroxide ^{-1} --> O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)[CH2] + O ^{-2}"
19670 17.280 18.890 30.045 -25.276 4.769 A + B --> AB "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + hydroxide ^{-1} --> O=N(=O)C1=CC(=[C](C(=[C]1)N(=O)=O)(C)O)N(=O)=O ^{-2}"
18871 251.127 248.795 246.579 -76.331 170.247 AB + C --> AC + B "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + [SH-] ^{-1} --> O=N(=O)c1[c]c(N(=O)=O)c(c([c]1)N(=O)=O)C + S ^{-2}"
18868 -5.352 -6.477 -9.495 -3.790 -13.285 AB + C --> AC + B "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + [SH-] ^{-1} --> O=N(=O)c1cc(S)c(c([c]1)N(=O)=O)C ^{-1} + O=[N]=O ^{-1}"
17231 -16.293 -17.068 -19.783 0.000 -19.783 AB + C --> AC + B "TNT theory{pspw} + hydroxide ^{-1} theory{pspw} --> O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} theory{pspw} + O theory{pspw}"
5633 255.015 255.590 253.611 -114.944 138.667 AB + C --> AC + B "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + hydroxide ^{-1} --> O=N(=O)c1[c]c(N(=O)=O)c(c([c]1)N(=O)=O)C + O ^{-2}"
5607 8.125 7.908 7.616 -8.920 -1.304 EA + BCD --> AB + CDE "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + O --> O=N(=O)c1cc(O)c(c([c]1)N(=O)=O)C ^{-1} + ON=O"
5167 17.269 19.006 31.044 -23.925 7.120 A + B --> AB "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + hydroxide ^{-1} --> O=N(=O)C1=CC(=[C](C(=[C]1)N(=O)=O)(C)O)N(=O)=O ^{-2}"
5091 20.237 22.462 34.196 -25.865 8.331 A + B --> AB "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + hydroxide ^{-1} --> O=N(=O)C1=C(C)C(=[CH](C(=[C]1)N(=O)=O)O)N(=O)=O ^{-2}"
5015 250.546 251.281 251.234 -115.625 135.608 AB + C --> AC + B "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + hydroxide ^{-1} --> O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)[CH2] + O ^{-2}"
3079 -44.076 -43.947 -45.409 16.538 -28.871 AB + C --> AC + B "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + hydroxide ^{-1} --> O=N(=O)c1cc(O)c(c([c]1)N(=O)=O)C ^{-1} + O=[N]=O ^{-1}"
1263 8.125 7.908 7.616 -8.920 -1.304 EA + BCD --> AB + CDE "O=N(=O)c1[c]c(N(=O)=O)c(c(c1)N(=O)=O)C ^{-1} + O --> O=N(=O)c1cc(O)c(c([c]1)N(=O)=O)C ^{-1} + ON=O"
All requests to Arrows were successful.
KEYWORDs -
reaction: :reaction
chinese_room: :chinese_room
molecule: :molecule
nmr: :nmr
predict: :predict
submitesmiles: :submitesmiles
nosubmitmissingesmiles
resubmitmissingesmiles
submitmachines: :submitmachines
useallentries
nomodelcorrect
eigenvalues: :eigenvalues
frequencies: :frequencies
nwoutput: :nwoutput
xyzfile: :xyzfile
alleigs: :alleigs
allfreqs: :allfreqs
reactionenumerate:
energytype:[erxn(gas) hrxn(gas) grxn(gas) delta_solvation grxn(aq)] :energytype
energytype:[kcal/mol kj/mol ev cm-1 ry hartree au] :energytype
tablereactions:
reaction: ... :reaction
reaction: ... :reaction
...
:tablereactions
tablemethods:
method: ... :method
method: ... :method
...
:tablemethods
:reactionenumerate
rotatebonds
xyzinput:
label: :label
xyzdata:
... xyz data ...
:xyzdata
:xyzinput
submitHf: :submitHf
nmrexp: :nmrexp
findreplace: old text | new text :findreplace
listnwjobs
fetchnwjob: :fetchnwjob
pushnwjob: :pushnwjob
printcsv: :printcsv
printeig: :printeig
printfreq: :printfreq
printxyz: :printxyz
printjobinfo: :printjobinfo
printnwout: :printnwout
badids: :badids
hup_string:
database:
table:
request_table:
listallesmiles
queuecheck
This software service and its documentation were developed at the Environmental Molecular Sciences Laboratory (EMSL) at Pacific Northwest National Laboratory, a multiprogram national laboratory, operated for the U.S. Department of Energy by Battelle under Contract Number DE-AC05-76RL01830. Support for this work was provided by the Department of Energy Office of Biological and Environmental Research, and Department of Defense environmental science and technology program (SERDP). THE SOFTWARE SERVICE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE SERVICE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE SERVICE.