 3D Periodic Editor 3D Molecular And Reaction Editor Expert Editor Microsoft Quantum Editor EMSL Aerosol Workshop Editor Manual
3D Periodic Editor 3D Molecular And Reaction Editor Expert Editor Microsoft Quantum Editor EMSL Aerosol Workshop Editor Manual Results from an EMSL Arrows Calculation
| EMSL Arrows is a revolutionary approach to materials and chemical simulations that uses NWChem and chemical computational databases to make materials and chemical modeling accessible via a broad spectrum of digital communications including posts to web APIs, social networks, and traditional email. |
Molecular modeling software has previously been extremely complex, making it prohibative to all but experts in the field, yet even experts can struggle to perform calculations. This service is designed to be used by experts and non-experts alike. Experts can carry out and keep track of large numbers of complex calculations with diverse levels of theories present in their workflows. Additionally, due to a streamlined and easy-to-use input, non-experts can carry out a wide variety of molecular modeling calculations previously not accessible to them.
The id(s) for emsiles = CC=C theory{dft} xc{b3lyp} basis{6-311++G(2d,2p)} solvation_type{COSMO} ^{0} are: 24181
Use id=% instead of esmiles to print other entries.
mformula = C3H6
iupac = prop-1-ene
PubChem = 8252
PubChem LCSS = 8252
cas = 115-07-1
kegg = C11505
synonyms = Propene; PROPYLENE; 1-Propene; Methylethylene; Methylethene; 1-Propylene; prop-1-ene; 115-07-1; Propene, pure; NCI-C50077; CH2=CH-CH3; n-propylene; CCRIS 1356; HSDB 175; 1-methylethylene; 2-methylethylene; CHEBI:16052; QQONPFPTGQHPMA-UHFFFAOYSA-N; R 1270; Hydrocarbons, C3; 2,2-propylene; propan-1,2-diyl; EINECS 204-062-1; UN1077; 90530-12-4; Polipropene 25; propane-1,2-diyl; Polypropylene, isotatic; 1-Propene, homopolymer; Polypropylene, isotactic; ACMC-1BBRS; AC1L1QLA; AC1Q2CAM; propylene, various grades; CH3CH=CH2; UNII-AUG1H506LY; AUG1H506LY; ETHENYL, 1-METHYL-; 182389_ALDRICH; 295663_ALDRICH; 427853_ALDRICH; 427861_ALDRICH; 427888_ALDRICH; 427896_ALDRICH; 428116_ALDRICH; 428175_ALDRICH; 428183_ALDRICH; 452149_ALDRICH; 452157_ALDRICH; 565628_ALDRICH; CHEMBL117213; 82222_FLUKA; CTK0H6486; 1-Propene, homopolymer, isotactic; Propene,ammoxidized,by-productsfrom; EINECS 292-050-7; AR-1C5550; c0067; LS-571; AKOS009156831; Propene, ammoxidized, by-products from; RL00588; UN 1077; Polypropylene, methyl-PSS nanoreinforced; OR079717; OR116117; OR205597; OR225679; OR238733; TC-165351; FT-0632493; InChI=1/C3H6/c1-3-2/h3H,1H2,2H; Propylene see also Petroleum gases, liquefied; 8241-EP2316824A1; C11505; R-1270; 10949-EP2269610A2; 10949-EP2270010A1; 10949-EP2270113A1; 10949-EP2272822A1; 10949-EP2272935A1; 10949-EP2272972A1; 10949-EP2272973A1; 10949-EP2275413A1; 10949-EP2275417A2; 10949-EP2275420A1; 10949-EP2277848A1; 10949-EP2277867A2; 10949-EP2277872A1; 10949-EP2277945A1; 10949-EP2280003A2; 10949-EP2281823A2; 10949-EP2284162A2; 10949-EP2284163A2; 10949-EP2286812A1; 10949-EP2287156A1; 10949-EP2289510A1; 10949-EP2289868A1; 10949-EP2289876A1; 10949-EP2289890A1; 10949-EP2292589A1; 10949-EP2292593A2; 10949-EP2292597A1; 10949-EP2292604A2; 10949-EP2295399A2; 10949-EP2295407A1; 10949-EP2295414A1; 10949-EP2298736A1; 10949-EP2298743A1; 10949-EP2298744A2; 10949-EP2298747A1; 10949-EP2298770A1; 10949-EP2298778A1; 10949-EP2298828A1; 10949-EP2299326A1; 10949-EP2301922A1; 10949-EP2301937A1; 10949-EP2305641A1; 10949-EP2305642A2; 10949-EP2305653A1; 10949-EP2305654A1; 10949-EP2305662A1; 10949-EP2305683A1; 10949-EP2308510A1; 10949-EP2308828A2; 10949-EP2308843A1; 10949-EP2308862A1; 10949-EP2308877A1; 10949-EP2311815A1; 10949-EP2311824A1; 10949-EP2311839A1; 10949-EP2314579A1; 10949-EP2314584A1; 10949-EP2314589A1; 10949-EP2316457A1; 10949-EP2316458A1; 10949-EP2316825A1; 10949-EP2316826A1; 10949-EP2316827A1; 10949-EP2316828A1; 10949-EP2316837A1; 10949-EP2370380A2; 10949-EP2371810A1; 10949-EP2372017A1; 10949-EP2374780A1; 10949-EP2374781A1; 10949-EP2374788A1; 10949-EP2377842A1; 10949-EP2380661A2; 10949-EP2380867A1; 16128-EP2281563A1; 16128-EP2308848A1; 16128-EP2316459A1; 17194-EP2272835A1; 17194-EP2272843A1; 17194-EP2272844A1; 17194-EP2281817A1; 17194-EP2314586A1; 27387-EP2275412A1; 27387-EP2275420A1; 27387-EP2292589A1; 27387-EP2295414A1; 27387-EP2295416A2; 27387-EP2295426A1; 27387-EP2295427A1; 27387-EP2298748A2; 27387-EP2298767A1; 27387-EP2305685A1; 27387-EP2305686A1; 27387-EP2308960A1; 27387-EP2311825A1; 27387-EP2311841A1; 27387-EP2314587A1; 92645-EP2277881A1; 92645-EP2289518A1; 92645-EP2292231A1; 92645-EP2305653A1; 92645-EP2305654A1; 92646-EP2292231A1; 92646-EP2305653A1; 100787-EP2301927A1; 100787-EP2374780A1; 147432-EP2289891A2; 147432-EP2292589A1; 1-Propene,ammoxidized,by-productsfrom,thermal-cracked; Propylene see also Petroleum gases, liquefied [UN1077] [Flammable gas]; Propylene see also Petroleum gases, liquefied [UN1077] [Flammable gas]; 33004-01-2; 676-63-1; 97102-85-7
Search Links to Other Online Resources (may not be available):
- Google Structure Search
- EPA CompTox Database
- Chemical Entities of Biological Interest (ChEBI)
- NIH ChemIDplus - A TOXNET DATABASE
- The Human Metabolome Database (HMDB)
- OECD eChemPortal
- Google Scholar
+==================================================+
|| Molecular Calculation ||
+==================================================+
Id = 24181
NWOutput = Link to NWChem Output (download)
Datafiles:
nwchemarrows.out-674215-2017-7-3-7:37:7 (download)
lumo-restricted.cube-674215-2017-7-3-7:37:7 (download)
homo-restricted.cube-674215-2017-7-3-7:37:7 (download)
nwchemarrows.out-335496-hrotor-2017-11-19-0:37:47 (download)
nwchemarrows.out-102448-hrotor-2017-11-24-17:37:28 (download)
mo_orbital_nwchemarrows.out-42274-2017-12-4-21:37:44 (download)
image_resset: api/image_reset/24181
Calculation performed by we29676
Numbers of cpus used for calculation = 4
Calculation walltime = 1139.800000 seconds (0 days 0 hours 18 minutes 59 seconds)
+----------------+
| Energetic Data |
+----------------+
Id = 24181
iupac = prop-1-ene
mformula = C3H6
inchi = InChI=1S/C3H6/c1-3-2/h3H,1H2,2H3
inchikey = QQONPFPTGQHPMA-UHFFFAOYSA-N
esmiles = CC=C theory{dft} xc{b3lyp} basis{6-311++G(2d,2p)} solvation_type{COSMO} ^{0}
calculation_type = ovc
theory = dft
xc = b3lyp
basis = 6-311++G(2d,2p)
charge,mult = 0 1
energy = -117.953032 Hartrees
enthalpy correct.= 0.084488 Hartrees
entropy = 63.192 cal/mol-K
solvation energy = -0.093 kcal/mol solvation_type = COSMO
Sitkoff cavity dispersion = 1.877 kcal/mol
Honig cavity dispersion = 5.083 kcal/mol
ASA solvent accesible surface area = 203.326 Angstrom2
ASA solvent accesible volume = 184.017 Angstrom3
+-----------------+
| Structural Data |
+-----------------+
JSmol: an open-source HTML5 viewer for chemical structures in 3D
Number of Atoms = 9
Units are Angstrom for bonds and degrees for angles
Type I J K L M Value
----------- ----- ----- ----- ----- ----- ----------
1 Stretch C1 C2 1.49815
2 Stretch C1 H4 1.09237
3 Stretch C1 H5 1.09309
4 Stretch C1 H6 1.08972
5 Stretch C2 C3 1.32780
6 Stretch C2 H7 1.08593
7 Stretch C3 H8 1.08336
8 Stretch C3 H9 1.08129
9 Bend C2 C1 H4 111.22836
10 Bend C2 C1 H5 110.81267
11 Bend C2 C1 H6 111.53988
12 Bend H4 C1 H5 106.62064
13 Bend H4 C1 H6 108.32886
14 Bend H5 C1 H6 108.12104
15 Bend C1 C2 C3 125.28903
16 Bend C1 C2 H7 116.02910
17 Bend C3 C2 H7 118.67960
18 Bend C2 C3 H8 121.56427
19 Bend C2 C3 H9 121.55854
20 Bend H8 C3 H9 116.87685
21 Dihedral C1 C2 C3 H8 0.46364
22 Dihedral C1 C2 C3 H9 -179.31649
23 Dihedral C3 C2 C1 H4 -126.58220
24 Dihedral C3 C2 C1 H5 114.99784
25 Dihedral C3 C2 C1 H6 -5.51676
26 Dihedral H4 C1 C2 H7 53.97584
27 Dihedral H5 C1 C2 H7 -64.44413
28 Dihedral H6 C1 C2 H7 175.04128
29 Dihedral H7 C2 C3 H8 179.89209
30 Dihedral H7 C2 C3 H9 0.11196
+-----------------+
| Eigenvalue Data |
+-----------------+
Id = 24181
iupac = prop-1-ene
mformula = C3H6
InChI = InChI=1S/C3H6/c1-3-2/h3H,1H2,2H3
smiles = CC=C
esmiles = CC=C theory{dft} xc{b3lyp} basis{6-311++G(2d,2p)} solvation_type{COSMO} ^{0}
theory = dft
xc = b3lyp
basis = 6-311++G(2d,2p)
charge = 0
mult = 1
solvation_type = COSMO
twirl webpage = TwirlMol Link
image webpage = GIF Image Link
Eigevalue Spectra
---------- 13.67 eV
----------
---- ----
----------
----------
----------
----------
----------
----------
----------
----------
---- ----
--- -- ---
-- -- -- -
---- ----
--- -- ---
---- ----
--- -- ---
---- ---- LUMO= -0.14 eV
HOMO= -7.38 eV ++++++++++
++++++++++
++++++++++
++++ ++++
++++++++++
++++++++++
++++++++++
-21.64 eV ++++++++++
spin eig occ ---------------------------- restricted -21.64 2.00 restricted -18.84 2.00 restricted -15.23 2.00 restricted -12.90 2.00 restricted -11.67 2.00 restricted -11.39 2.00 restricted -10.28 2.00 restricted -9.55 2.00 restricted -7.38 2.00 restricted -0.14 0.00 restricted -0.12 0.00 restricted 0.46 0.00 restricted 0.52 0.00 restricted 0.95 0.00 restricted 1.26 0.00 restricted 1.55 0.00 restricted 2.05 0.00 restricted 2.27 0.00 restricted 2.45 0.00 restricted 2.70 0.00 restricted 3.07 0.00 restricted 3.30 0.00 restricted 3.66 0.00 restricted 3.78 0.00 restricted 3.88 0.00 restricted 3.96 0.00 restricted 4.38 0.00 restricted 4.56 0.00 restricted 5.12 0.00 restricted 5.37 0.00 restricted 5.70 0.00 restricted 6.11 0.00 restricted 7.07 0.00 restricted 7.92 0.00 restricted 9.57 0.00 restricted 10.20 0.00 restricted 10.95 0.00 restricted 11.73 0.00 restricted 11.93 0.00 restricted 12.38 0.00 restricted 12.91 0.00 restricted 13.67 0.00
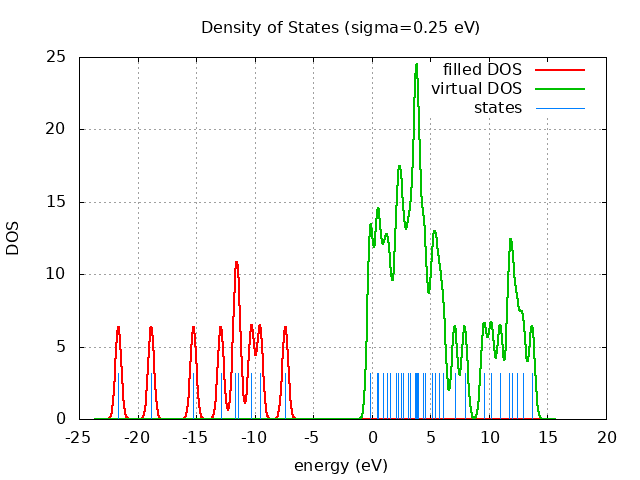
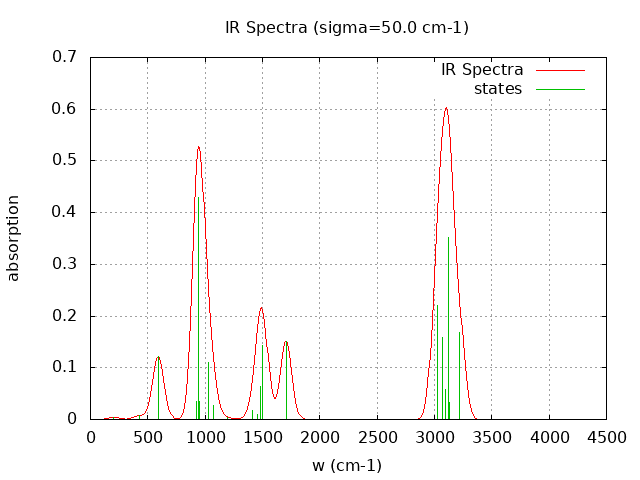
+----------------------------------------+ | Vibrational Density of States Analysis | +----------------------------------------+ Total number of frequencies = 27 Total number of negative frequencies = 0 Number of lowest frequencies = 2 (frequency threshold = 500 ) Exact dos norm = 21.000 Generating vibrational DOS Generating model vibrational DOS to have a proper norm 10.00 21.00 2.00 21.00 50.00 21.00 2.00 21.00 100.00 20.98 1.98 21.00 Generating IR Spectra Writing vibrational density of states (DOS) to vdos.dat Writing model vibrational density of states (DOS_FIXED) to vdos-model.dat Writing IR spectra to irdos.dat Temperature= 298.15 zero-point correction to energy = 49.861 kcal/mol ( 0.079459) vibrational contribution to enthalpy correction = 50.648 kcal/mol ( 0.080713) vibrational contribution to Entropy = 4.062 cal/mol-k hindered rotor enthalpy correction = 0.077 kcal/mol ( 0.000122) hindered rotor entropy correction = 0.346 cal/mol-k

DOS sigma = 10.000000
- vibrational DOS enthalpy correction = 0.080714 kcal/mol ( 50.649 kcal/mol)
- model vibrational DOS enthalpy correction = 0.080713 kcal/mol ( 50.648 kcal/mol)
- vibrational DOS Entropy = 0.000006 ( 4.067 cal/mol-k)
- model vibrational DOS Entropy = 0.000006 ( 4.065 cal/mol-k)
- original gas Energy = -117.953032 (-74016.644 kcal/mol)
- original gas Enthalpy = -117.868544 (-73963.627 kcal/mol, delta= 0.000)
- unajusted DOS gas Enthalpy = -117.868543 (-73963.627 kcal/mol, delta= 0.000)
- model DOS gas Enthalpy = -117.868544 (-73963.627 kcal/mol, delta= -0.000)
- original gas Entropy = 0.000101 ( 63.192 cal/mol-k,delta= 0.000)
- unajusted DOS gas Entropy = 0.000101 ( 63.196 cal/mol-k,delta= 0.004)
- model DOS gas Entropy = 0.000101 ( 63.195 cal/mol-k,delta= 0.003)
- original gas Free Energy = -117.898568 (-73982.468 kcal/mol, delta= 0.000)
- unadjusted DOS gas Free Energy = -117.898569 (-73982.469 kcal/mol, delta= -0.001)
- model DOS gas Free Energy = -117.898570 (-73982.469 kcal/mol, delta= -0.001)
- original sol Free Energy = -117.898717 (-73982.561 kcal/mol)
- unadjusted DOS sol Free Energy = -117.898718 (-73982.562 kcal/mol)
- model DOS sol Free Energy = -117.898719 (-73982.562 kcal/mol)
DOS sigma = 10.000000 - Estimates including Hindered Rotor Corrections
- vibrational DOS enthalpy correction = 0.080714 kcal/mol ( 50.649 kcal/mol)
- model vibrational DOS enthalpy correction = 0.080713 kcal/mol ( 50.648 kcal/mol)
- vibrational DOS Entropy = 0.000006 ( 4.067 cal/mol-k)
- model vibrational DOS Entropy = 0.000006 ( 4.065 cal/mol-k)
- original gas Energy = -117.953032 (-74016.644 kcal/mol)
- original gas Enthalpy = -117.868421 (-73963.551 kcal/mol, delta= 0.000)
- unajusted DOS gas Enthalpy = -117.868421 (-73963.550 kcal/mol, delta= 0.000)
- model DOS gas Enthalpy = -117.868422 (-73963.551 kcal/mol, delta= -0.000)
- original gas Entropy = 0.000101 ( 63.538 cal/mol-k,delta= 0.000)
- unajusted DOS gas Entropy = 0.000101 ( 63.542 cal/mol-k,delta= 0.004)
- model DOS gas Entropy = 0.000101 ( 63.540 cal/mol-k,delta= 0.003)
- original gas Free Energy = -117.898610 (-73982.494 kcal/mol, delta= 0.000)
- unadjusted DOS gas Free Energy = -117.898612 (-73982.495 kcal/mol, delta= -0.001)
- model DOS gas Free Energy = -117.898612 (-73982.495 kcal/mol, delta= -0.001)
- original sol Free Energy = -117.898759 (-73982.588 kcal/mol)
- unadjusted DOS sol Free Energy = -117.898760 (-73982.589 kcal/mol)
- model DOS sol Free Energy = -117.898761 (-73982.589 kcal/mol)
DOS sigma = 50.000000
- vibrational DOS enthalpy correction = 0.080727 kcal/mol ( 50.657 kcal/mol)
- model vibrational DOS enthalpy correction = 0.080727 kcal/mol ( 50.657 kcal/mol)
- vibrational DOS Entropy = 0.000007 ( 4.171 cal/mol-k)
- model vibrational DOS Entropy = 0.000007 ( 4.171 cal/mol-k)
- original gas Energy = -117.953032 (-74016.644 kcal/mol)
- original gas Enthalpy = -117.868544 (-73963.627 kcal/mol, delta= 0.000)
- unajusted DOS gas Enthalpy = -117.868530 (-73963.618 kcal/mol, delta= 0.009)
- model DOS gas Enthalpy = -117.868530 (-73963.618 kcal/mol, delta= 0.009)
- original gas Entropy = 0.000101 ( 63.192 cal/mol-k,delta= 0.000)
- unajusted DOS gas Entropy = 0.000101 ( 63.300 cal/mol-k,delta= 0.108)
- model DOS gas Entropy = 0.000101 ( 63.300 cal/mol-k,delta= 0.108)
- original gas Free Energy = -117.898568 (-73982.468 kcal/mol, delta= 0.000)
- unadjusted DOS gas Free Energy = -117.898606 (-73982.491 kcal/mol, delta= -0.024)
- model DOS gas Free Energy = -117.898606 (-73982.491 kcal/mol, delta= -0.024)
- original sol Free Energy = -117.898717 (-73982.561 kcal/mol)
- unadjusted DOS sol Free Energy = -117.898754 (-73982.585 kcal/mol)
- model DOS sol Free Energy = -117.898754 (-73982.585 kcal/mol)
DOS sigma = 50.000000 - Estimates including Hindered Rotor Corrections
- vibrational DOS enthalpy correction = 0.080727 kcal/mol ( 50.657 kcal/mol)
- model vibrational DOS enthalpy correction = 0.080727 kcal/mol ( 50.657 kcal/mol)
- vibrational DOS Entropy = 0.000007 ( 4.171 cal/mol-k)
- model vibrational DOS Entropy = 0.000007 ( 4.171 cal/mol-k)
- original gas Energy = -117.953032 (-74016.644 kcal/mol)
- original gas Enthalpy = -117.868421 (-73963.551 kcal/mol, delta= 0.000)
- unajusted DOS gas Enthalpy = -117.868408 (-73963.542 kcal/mol, delta= 0.009)
- model DOS gas Enthalpy = -117.868408 (-73963.542 kcal/mol, delta= 0.009)
- original gas Entropy = 0.000101 ( 63.538 cal/mol-k,delta= 0.000)
- unajusted DOS gas Entropy = 0.000101 ( 63.646 cal/mol-k,delta= 0.108)
- model DOS gas Entropy = 0.000101 ( 63.646 cal/mol-k,delta= 0.108)
- original gas Free Energy = -117.898610 (-73982.494 kcal/mol, delta= 0.000)
- unadjusted DOS gas Free Energy = -117.898648 (-73982.518 kcal/mol, delta= -0.024)
- model DOS gas Free Energy = -117.898648 (-73982.518 kcal/mol, delta= -0.024)
- original sol Free Energy = -117.898759 (-73982.588 kcal/mol)
- unadjusted DOS sol Free Energy = -117.898797 (-73982.611 kcal/mol)
- model DOS sol Free Energy = -117.898797 (-73982.611 kcal/mol)
DOS sigma = 100.000000
- vibrational DOS enthalpy correction = 0.080751 kcal/mol ( 50.672 kcal/mol)
- model vibrational DOS enthalpy correction = 0.080775 kcal/mol ( 50.687 kcal/mol)
- vibrational DOS Entropy = 0.000007 ( 4.483 cal/mol-k)
- model vibrational DOS Entropy = 0.000007 ( 4.518 cal/mol-k)
- original gas Energy = -117.953032 (-74016.644 kcal/mol)
- original gas Enthalpy = -117.868544 (-73963.627 kcal/mol, delta= 0.000)
- unajusted DOS gas Enthalpy = -117.868505 (-73963.603 kcal/mol, delta= 0.024)
- model DOS gas Enthalpy = -117.868481 (-73963.588 kcal/mol, delta= 0.039)
- original gas Entropy = 0.000101 ( 63.192 cal/mol-k,delta= 0.000)
- unajusted DOS gas Entropy = 0.000101 ( 63.613 cal/mol-k,delta= 0.421)
- model DOS gas Entropy = 0.000101 ( 63.647 cal/mol-k,delta= 0.455)
- original gas Free Energy = -117.898568 (-73982.468 kcal/mol, delta= 0.000)
- unadjusted DOS gas Free Energy = -117.898730 (-73982.569 kcal/mol, delta= -0.101)
- model DOS gas Free Energy = -117.898722 (-73982.565 kcal/mol, delta= -0.097)
- original sol Free Energy = -117.898717 (-73982.561 kcal/mol)
- unadjusted DOS sol Free Energy = -117.898879 (-73982.663 kcal/mol)
- model DOS sol Free Energy = -117.898871 (-73982.658 kcal/mol)
DOS sigma = 100.000000 - Estimates including Hindered Rotor Corrections
- vibrational DOS enthalpy correction = 0.080751 kcal/mol ( 50.672 kcal/mol)
- model vibrational DOS enthalpy correction = 0.080775 kcal/mol ( 50.687 kcal/mol)
- vibrational DOS Entropy = 0.000007 ( 4.483 cal/mol-k)
- model vibrational DOS Entropy = 0.000007 ( 4.518 cal/mol-k)
- original gas Energy = -117.953032 (-74016.644 kcal/mol)
- original gas Enthalpy = -117.868421 (-73963.551 kcal/mol, delta= 0.000)
- unajusted DOS gas Enthalpy = -117.868383 (-73963.527 kcal/mol, delta= 0.024)
- model DOS gas Enthalpy = -117.868359 (-73963.512 kcal/mol, delta= 0.039)
- original gas Entropy = 0.000101 ( 63.538 cal/mol-k,delta= 0.000)
- unajusted DOS gas Entropy = 0.000102 ( 63.959 cal/mol-k,delta= 0.421)
- model DOS gas Entropy = 0.000102 ( 63.993 cal/mol-k,delta= 0.455)
- original gas Free Energy = -117.898610 (-73982.494 kcal/mol, delta= 0.000)
- unadjusted DOS gas Free Energy = -117.898772 (-73982.596 kcal/mol, delta= -0.101)
- model DOS gas Free Energy = -117.898764 (-73982.591 kcal/mol, delta= -0.097)
- original sol Free Energy = -117.898759 (-73982.588 kcal/mol)
- unadjusted DOS sol Free Energy = -117.898921 (-73982.689 kcal/mol)
- model DOS sol Free Energy = -117.898913 (-73982.684 kcal/mol)
Normal Mode Frequency (cm-1) IR Intensity (arbitrary)
----------- ---------------- ------------------------
1 -0.000 0.121
2 -0.000 0.093
3 0.000 0.091
4 0.000 0.058
5 0.000 0.021
6 0.000 0.152
7 204.060 0.537
8 428.060 0.907
9 591.570 15.243
10 921.490 4.449
11 943.180 53.807
12 953.010 4.482
13 1025.190 13.922
14 1074.310 3.387
15 1194.160 0.403
16 1335.650 0.017
17 1414.340 2.173
18 1456.650 1.237
19 1484.800 7.973
20 1500.600 17.813
21 1706.000 19.009
22 3023.270 27.630
23 3066.120 19.865
24 3099.850 7.213
25 3124.680 44.174
26 3133.630 4.215
27 3214.380 21.008
@@@@@@@@@@@@@@@@@@@@@@ START ROTOR @@@@@@@@@@@@@@@@@@@@@@@@@@

rotation xyz movie
Hindered Rotor Calculation:
Temperature = 298.15
rbond = 2 1
rgroup = 4 5 6
rsym_num = 3
nphi = 72
NNmax = 2000
Total inertia:
I=
1.268558e+05 1.089816e+05 -3.523893e+04
1.089816e+05 3.012324e+05 1.795887e+04
-3.523893e+04 1.795887e+04 3.983894e+05
IA = 6.972073e+04
IB = 3.535936e+05
IC = 4.031632e+05
VA = <-8.946862e-01 4.304294e-01 -1.194450e-01>
VB = <4.354418e-01 9.000312e-01 -1.828382e-02>
VC = <-9.963429e-02 6.836961e-02 9.926725e-01>
Group inertia:
Isub=
2.029202e+04 -2.589279e+02 9.562406e+02
-2.589279e+02 1.008056e+04 -1.989457e+01
9.562406e+02 -1.989457e+01 1.040786e+04
rotation axis bt = -9.948275e-01 1.999861e-02 -9.959055e-02
Im0 = bt*Isub*b = 2.038976e+04
bt*b = 1.000000e+00
Free Rotation, Pitzer-Gwinn Formula:
T = 298.150000
Im = 15261.182076 (3.915129e-47 Kg-m2)
n = 3
Qf = 3.178081
Uf = 0.296063 kcal/mol (0.000472 au)
Sf = 3.289367 cal/mol-K
Free Rotation, Cannonical Formula, 5001 eigenvalues used:
T = 298.150 K
sigma = 3.000000
Im = 15261.182076
Qf = 9.512195
dQf/dT = 0.015952 1/K
Uf = 0.296063 kcal/mol ( 0.000472 au)
Sf = 3.284769 cal/mol-K
IA = 69720.730391 (1.788627e-46 Kg-m2)
IB = 353593.639168 (9.071150e-46 Kg-m2)
IC = 403163.231130 (1.034282e-45 Kg-m2)
theory = dft
drion/dphi = 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 -0.037355
drion/dphi = 0.858909 0.545624 -0.067280 -0.892511 0.492849 0.101430 0.024814 -1.008221 0.000000 0.000000
drion/dphi = 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000
tphi = 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 0.036028
tphi = -0.828402 -0.526244 0.064444 0.854893 -0.472077 -0.098724 -0.024152 0.981319 0.000000 0.000000
tphi = 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000 0.000000
Cannonical Formula Results for Hindered Rotation:
Nmax = 1152
T = 298.150 K
sigma = 3.000000
Im = 15261.182076
Qf = 3.060813
dQf/dT = 0.012412 1/K
Uf = 0.715889 kcal/mol ( 0.001141 au)
Sf = 2.440960 cal/mol-K
Uvib(bad)= 0.639303 kcal/mol ( 0.001019 au)
Svib(bad)= 2.095233 cal/mol-K ( 0.000003 au)
Ucorrect = 0.076587 kcal/mol ( 0.000122 au)
Scorrect = 0.345728 cal/mol-K ( 0.000001 au)
Frequency Overlap Results for Hindered Rotation:
0 w=-0.000 drphit*freq=-0.108 weight=0.000
1 w=-0.000 drphit*freq=0.000 weight=0.000
2 w=0.000 drphit*freq=0.000 weight=0.000
3 w=0.000 drphit*freq=0.008 weight=0.000
4 w=0.000 drphit*freq=0.063 weight=0.000
5 w=0.000 drphit*freq=0.000 weight=0.000
6 w=204.060 drphit*freq=-0.434 weight=1.000
7 w=428.060 drphit*freq=0.000 weight=0.000
8 w=591.570 drphit*freq=0.000 weight=0.000
9 w=921.490 drphit*freq=0.000 weight=0.000
10 w=943.180 drphit*freq=0.000 weight=0.000
11 w=953.010 drphit*freq=0.000 weight=0.000
12 w=1025.190 drphit*freq=0.000 weight=0.000
13 w=1074.310 drphit*freq=0.000 weight=0.000
14 w=1194.160 drphit*freq=0.000 weight=0.000
15 w=1335.650 drphit*freq=0.000 weight=0.000
16 w=1414.340 drphit*freq=0.000 weight=0.000
17 w=1456.650 drphit*freq=0.000 weight=0.000
18 w=1484.800 drphit*freq=0.000 weight=0.000
19 w=1500.600 drphit*freq=0.000 weight=0.000
20 w=1706.000 drphit*freq=0.000 weight=0.000
21 w=3023.270 drphit*freq=0.000 weight=0.000
22 w=3066.120 drphit*freq=0.000 weight=0.000
23 w=3099.850 drphit*freq=0.000 weight=0.000
24 w=3124.680 drphit*freq=0.000 weight=0.000
25 w=3133.630 drphit*freq=0.000 weight=0.000
26 w=3214.380 drphit*freq=0.000 weight=0.000
hindered potential = -117.953032 -117.952867 -117.952607 -117.952268 -117.951874 -117.951450 -117.951026 -117.950629 -117.950288 -117.950020 -117.949850 -117.949786 -117.949834 -117.949991 -117.950240 -117.950567 -117.950949 -117.951361 -117.951773 -117.952157 -117.952487 -117.952740 -117.952899 -117.952954 -117.952902 -117.952745 -117.952495 -117.952168 -117.951786 -117.951377 -117.950969 -117.950589 -117.950264 -117.950013 -117.949860 -117.949812 -117.949874 -117.950040 -117.950302 -117.950636 -117.951023 -117.951435 -117.951844 -117.952223 -117.952544 -117.952787 -117.952934 -117.952976 -117.952910 -117.952741 -117.952479 -117.952141 -117.951751 -117.951336 -117.950923 -117.950542 -117.950218 -117.949974 -117.949826 -117.949786 -117.949856 -117.950032 -117.950302 -117.950649 -117.951048 -117.951474 -117.951897 -117.952289 -117.952624 -117.952879 -117.953038 -117.953089
@@@@@@@@@@@@@@@@@@@@@@ END ROTOR @@@@@@@@@@@@@@@@@@@@@@@@@@
+-------------------------------------+
| Reactions Contained in the Database |
+-------------------------------------+
Reactions Containing INCHIKEY = QQONPFPTGQHPMA-UHFFFAOYSA-N
Reactionid Erxn(gas) Hrxn(gas) Grxn(gas) delta_Solv Grxn(aq) ReactionType Reaction
21050 -256.195 -255.758 -255.247 89.655 -66.992 AB + C --> AC + B "C[C]=C mult{2} xc{b3lyp} + [OH3+] xc{b3lyp} + [SHE] xc{b3lyp} --> CC=C xc{b3lyp} + water xc{b3lyp}"
20971 -235.703 -235.461 -235.417 89.140 -47.677 AB + C --> AC + B "C=C[CH2] mult{2} xc{b3lyp} + [OH3+] xc{b3lyp} + [SHE] xc{b3lyp} --> CC=C xc{b3lyp} + water xc{b3lyp}"
20966 3.563 3.611 3.788 0.023 3.811 ABC + DE --> DBE + AC "CC=C xc{b3lyp} + CC xc{b3lyp} --> CCC xc{b3lyp} + C=C xc{b3lyp}"
20965 3.563 3.611 3.788 0.023 3.811 ABC + DE --> DBE + AC "CC=C xc{b3lyp} + CC xc{b3lyp} --> CCC xc{b3lyp} + C=C xc{b3lyp}"
20964 -254.900 -254.432 -253.957 89.236 -66.121 AB + C --> AC + B "C[C]=C xc{m06-2x} + [OH3+] xc{m06-2x} + [SHE] xc{m06-2x} --> CC=C xc{m06-2x} + water xc{m06-2x}"
20860 -34.923 -27.867 -18.873 -0.022 -18.895 AB + CD --> CABD "CC=C + [3H][3H] --> CCC"
20831 -425.926 -418.631 -410.972 258.068 -54.304 A + B --> AB "C[C]=C xc{m06-2x} + [H+] xc{m06-2x} + [SHE] xc{m06-2x} --> CC=C xc{m06-2x}"
20748 -431.521 -424.326 -416.708 258.348 -59.760 A + B --> AB "CC=[CH] xc{b3lyp} + [H+] xc{b3lyp} + [SHE] xc{b3lyp} --> CC=C xc{b3lyp}"
20743 -68.454 -68.347 -76.437 -78.097 -55.934 AB --> A + B "[CH2]C(C)Cl mult{2} xc{b3lyp} + [SHE] xc{b3lyp} --> CC=C xc{b3lyp} + [Cl-] xc{b3lyp}"
20742 -68.454 -68.347 -76.437 -78.097 -55.934 AB --> A + B "[CH2]C(C)Cl mult{2} xc{b3lyp} + [SHE] xc{b3lyp} --> CC=C xc{b3lyp} + [Cl-] xc{b3lyp}"
20680 -406.460 -399.415 -392.189 257.762 -35.827 A + B --> AB "C=C[CH2] xc{b3lyp} + [H+] xc{b3lyp} + [SHE] xc{b3lyp} --> CC=C xc{b3lyp}"
20637 -72.199 -72.273 -80.099 -78.751 -60.250 ABCD --> BCA + D "[CH2]CCCl mult{2} xc{b3lyp} + [SHE] xc{b3lyp} --> CC=C xc{b3lyp} + [Cl-] xc{b3lyp}"
20630 47.448 40.421 31.311 -1.642 29.669 CABD --> AB + CD "C=CC --> C#CC + [H][H]"
20617 -59.426 -59.343 -67.508 -78.932 -47.839 AB --> A + B "[CH2]C(C)Cl mult{2} xc{pbe0} + [SHE] xc{pbe0} --> CC=C xc{pbe0} + [Cl-] xc{pbe0}"
20616 -59.426 -59.343 -67.508 -78.932 -47.839 AB --> A + B "[CH2]C(C)Cl mult{2} xc{pbe0} + [SHE] xc{pbe0} --> CC=C xc{pbe0} + [Cl-] xc{pbe0}"
20559 -426.951 -419.713 -412.018 258.277 -55.141 A + B --> AB "C[C]=C xc{b3lyp} + [H+] xc{b3lyp} + [SHE] xc{b3lyp} --> CC=C xc{b3lyp}"
20555 -260.764 -260.371 -259.937 89.726 -71.611 AB + C --> AC + B "CC=[CH] xc{b3lyp} + [OH3+] xc{b3lyp} + [SHE] xc{b3lyp} --> CC=C xc{b3lyp} + water xc{b3lyp}"
20425 -47.310 -48.009 -57.386 12.316 -45.070 ABCD + E --> A + BC + DE "CCCCl + [OH-] --> C=CC + O + [Cl-]"
19921 -46.040 -46.654 -56.218 6.669 -49.549 ABCD + E --> A + BC + DE "CCCCl xc{m06-2x} + [OH-] xc{m06-2x} --> C=CC xc{m06-2x} + O xc{m06-2x} + [Cl-] xc{m06-2x}"
17400 -43.337 -44.069 -53.479 12.326 -41.153 ABCD + E --> A + BC + DE "CCCCl theory{ccsd(t)} + [OH-] theory{ccsd(t)} --> C=CC theory{ccsd(t)} + O theory{ccsd(t)} + [Cl-] theory{ccsd(t)}"
16748 47.408 40.411 31.340 -1.602 29.738 CABD --> AB + CD "C=CC --> C#CC + [H][H]"
14771 -59.823 -59.746 -67.875 -78.137 -47.412 AB --> A + B "[CH2]C(C)Cl mult{2} theory{ccsd(t)} + [SHE] theory{ccsd(t)} --> CC=C theory{ccsd(t)} + [Cl-] theory{ccsd(t)}"
14770 -59.823 -59.746 -67.875 -78.137 -47.412 AB --> A + B "[CH2]C(C)Cl mult{2} theory{ccsd(t)} + [SHE] theory{ccsd(t)} --> CC=C theory{ccsd(t)} + [Cl-] theory{ccsd(t)}"
7861 -41.169 -41.849 -51.170 11.768 -39.403 ABCD + E --> A + BC + DE "CCCCl xc{pbe} + [OH-] xc{pbe} --> C=CC xc{pbe} + O xc{pbe} + [Cl-] xc{pbe}"
7860 -257.231 -256.945 -256.444 90.036 -67.808 AB + C --> AC + B "CC=[CH] mult{2} xc{pbe} + [OH3+] xc{pbe} + [SHE] xc{pbe} --> CC=C xc{pbe} + water xc{pbe}"
7858 -252.280 -251.903 -251.276 90.006 -62.670 AB + C --> AC + B "C[C]=C mult{2} xc{pbe} + [OH3+] xc{pbe} + [SHE] xc{pbe} --> CC=C xc{pbe} + water xc{pbe}"
7855 -232.971 -232.873 -232.814 89.389 -44.825 AB + C --> AC + B "C=C[CH2] mult{2} xc{pbe} + [OH3+] xc{pbe} + [SHE] xc{pbe} --> CC=C xc{pbe} + water xc{pbe}"
7747 -256.532 -256.291 -255.877 89.707 -67.570 AB + C --> AC + B "CC=[CH] mult{2} xc{pbe0} + [OH3+] xc{pbe0} + [SHE] xc{pbe0} --> CC=C xc{pbe0} + water xc{pbe0}"
7555 -232.161 -231.972 -231.920 89.161 -44.159 AB + C --> AC + B "C=C[CH2] xc{pbe0} + [OH3+] xc{pbe0} + [SHE] xc{pbe0} --> CC=C xc{pbe0} + water xc{pbe0}"
7359 -251.976 -251.677 -251.183 89.666 -62.917 AB + C --> AC + B "C[C]=C xc{pbe0} + [OH3+] xc{pbe0} + [SHE] xc{pbe0} --> CC=C xc{pbe0} + water xc{pbe0}"
7302 3.902 3.940 4.137 0.125 4.261 ABC + DE --> DBE + AC "CC=C xc{pbe} + CC xc{pbe} --> CCC xc{pbe} + C=C xc{pbe}"
7301 3.902 3.940 4.137 0.125 4.261 ABC + DE --> DBE + AC "CC=C xc{pbe} + CC xc{pbe} --> CCC xc{pbe} + C=C xc{pbe}"
7271 2.974 2.717 2.836 0.083 2.919 ABC + DE --> DBE + AC "CC=C xc{m06-2x} + CC xc{m06-2x} --> CCC xc{m06-2x} + C=C xc{m06-2x}"
7270 2.974 2.717 2.836 0.083 2.919 ABC + DE --> DBE + AC "CC=C xc{m06-2x} + CC xc{m06-2x} --> CCC xc{m06-2x} + C=C xc{m06-2x}"
7023 -236.328 -235.710 -235.621 88.949 -48.073 AB + C --> AC + B "C=C[CH2] xc{m06-2x} + [OH3+] xc{m06-2x} + [SHE] xc{m06-2x} --> CC=C xc{m06-2x} + water xc{m06-2x}"
6800 -258.744 -258.313 -257.933 89.316 -70.017 AB + C --> AC + B "CC=[CH] xc{m06-2x} + [OH3+] xc{m06-2x} + [SHE] xc{m06-2x} --> CC=C xc{m06-2x} + water xc{m06-2x}"
5658 -254.900 -254.432 -253.957 89.286 -66.070 AB + C --> AC + B "C[C]=C xc{m06-2x} + [OH3+] xc{m06-2x} + [SHE] xc{m06-2x} --> CC=C xc{m06-2x} + water xc{m06-2x}"
2678 3.691 3.727 3.899 0.101 4.000 ABC + DE --> DBE + AC "CC=C xc{pbe0} + CC xc{pbe0} --> CCC xc{pbe0} + C=C xc{pbe0}"
2677 3.691 3.727 3.899 0.101 4.000 ABC + DE --> DBE + AC "CC=C xc{pbe0} + CC xc{pbe0} --> CCC xc{pbe0} + C=C xc{pbe0}"
2675 2.924 3.025 3.243 0.055 3.298 ABC + DE --> DBE + AC "CC=C theory{mp2} + CC theory{mp2} --> CCC theory{mp2} + C=C theory{mp2}"
2674 2.924 3.025 3.243 0.055 3.298 ABC + DE --> DBE + AC "CC=C theory{mp2} + CC theory{mp2} --> CCC theory{mp2} + C=C theory{mp2}"
2667 3.523 3.601 3.817 0.064 3.881 ABC + DE --> DBE + AC "CC=C xc{b3lyp} + CC xc{b3lyp} --> CCC xc{b3lyp} + C=C xc{b3lyp}"
2666 3.523 3.601 3.817 0.064 3.881 ABC + DE --> DBE + AC "CC=C xc{b3lyp} + CC xc{b3lyp} --> CCC xc{b3lyp} + C=C xc{b3lyp}"
2662 2.974 2.704 2.821 0.094 2.915 ABC + DE --> DBE + AC "CC=C xc{m06-2x} + CC xc{m06-2x} --> CCC xc{m06-2x} + C=C xc{m06-2x}"
2661 2.974 2.704 2.821 0.094 2.915 ABC + DE --> DBE + AC "CC=C xc{m06-2x} + CC xc{m06-2x} --> CCC xc{m06-2x} + C=C xc{m06-2x}"
2659 3.901 3.932 4.129 0.144 4.273 ABC + DE --> DBE + AC "CC=C xc{pbe} + CC xc{pbe} --> CCC xc{pbe} + C=C xc{pbe}"
2658 3.901 3.932 4.129 0.144 4.273 ABC + DE --> DBE + AC "CC=C xc{pbe} + CC xc{pbe} --> CCC xc{pbe} + C=C xc{pbe}"
2031 -253.980 -253.573 -253.100 89.665 -64.836 AB + C --> AC + B "C[C]=C mult{2} theory{ccsd(t)} + [OH3+] theory{ccsd(t)} + [SHE] theory{ccsd(t)} --> CC=C theory{ccsd(t)} + water theory{ccsd(t)}"
2030 -257.147 -256.783 -256.388 89.736 -68.052 AB + C --> AC + B "CC=[CH] mult{2} theory{ccsd(t)} + [OH3+] theory{ccsd(t)} + [SHE] theory{ccsd(t)} --> CC=C theory{ccsd(t)} + water theory{ccsd(t)}"
2027 -426.021 -418.813 -411.157 258.237 -54.320 A + B --> AB "C[C]=C mult{2} theory{ccsd(t)} + [H+] theory{ccsd(t)} + [SHE] theory{ccsd(t)} --> CC=C theory{ccsd(t)}"
2024 -429.188 -422.023 -414.445 258.308 -57.537 A + B --> AB "CC=[CH] mult{2} theory{ccsd(t)} + [H+] theory{ccsd(t)} + [SHE] theory{ccsd(t)} --> CC=C theory{ccsd(t)}"
1940 -236.328 -235.723 -235.634 88.949 -48.085 AB + C --> AC + B "C=C[CH2] mult{2} xc{m06-2x} + [OH3+] xc{m06-2x} + [SHE] xc{m06-2x} --> CC=C xc{m06-2x} + water xc{m06-2x}"
1939 -232.971 -232.878 -232.818 89.440 -44.779 AB + C --> AC + B "C=C[CH2] mult{2} xc{pbe} + [OH3+] xc{pbe} + [SHE] xc{pbe} --> CC=C xc{pbe} + water xc{pbe}"
1938 -232.161 -231.983 -231.930 89.201 -44.129 AB + C --> AC + B "C=C[CH2] mult{2} xc{pbe0} + [OH3+] xc{pbe0} + [SHE] xc{pbe0} --> CC=C xc{pbe0} + water xc{pbe0}"
1937 -235.664 -235.462 -235.462 89.120 -47.742 AB + C --> AC + B "C=C[CH2] mult{2} xc{b3lyp} + [OH3+] xc{b3lyp} + [SHE] xc{b3lyp} --> CC=C xc{b3lyp} + water xc{b3lyp}"
1930 -254.900 -254.445 -253.970 89.286 -66.083 AB + C --> AC + B "C[C]=C mult{2} xc{m06-2x} + [OH3+] xc{m06-2x} + [SHE] xc{m06-2x} --> CC=C xc{m06-2x} + water xc{m06-2x}"
1929 -252.280 -251.907 -251.280 90.056 -62.624 AB + C --> AC + B "C[C]=C mult{2} xc{pbe} + [OH3+] xc{pbe} + [SHE] xc{pbe} --> CC=C xc{pbe} + water xc{pbe}"
1928 -251.976 -251.687 -251.194 89.706 -62.887 AB + C --> AC + B "C[C]=C mult{2} xc{pbe0} + [OH3+] xc{pbe0} + [SHE] xc{pbe0} --> CC=C xc{pbe0} + water xc{pbe0}"
1927 -256.155 -255.749 -255.276 89.615 -67.062 AB + C --> AC + B "C[C]=C mult{2} xc{b3lyp} + [OH3+] xc{b3lyp} + [SHE] xc{b3lyp} --> CC=C xc{b3lyp} + water xc{b3lyp}"
1926 -258.744 -258.326 -257.946 89.316 -70.030 AB + C --> AC + B "CC=[CH] mult{2} xc{m06-2x} + [OH3+] xc{m06-2x} + [SHE] xc{m06-2x} --> CC=C xc{m06-2x} + water xc{m06-2x}"
1925 -257.231 -256.949 -256.449 90.086 -67.762 AB + C --> AC + B "CC=[CH] mult{2} xc{pbe} + [OH3+] xc{pbe} + [SHE] xc{pbe} --> CC=C xc{pbe} + water xc{pbe}"
1924 -256.532 -256.302 -255.888 89.748 -67.541 AB + C --> AC + B "CC=[CH] mult{2} xc{pbe0} + [OH3+] xc{pbe0} + [SHE] xc{pbe0} --> CC=C xc{pbe0} + water xc{pbe0}"
1923 -260.724 -260.361 -259.966 89.686 -71.680 AB + C --> AC + B "CC=[CH] mult{2} xc{b3lyp} + [OH3+] xc{b3lyp} + [SHE] xc{b3lyp} --> CC=C xc{b3lyp} + water xc{b3lyp}"
1913 -234.187 -233.985 -233.985 89.170 -46.215 AB + C --> AC + B "C=C[CH2] mult{2} theory{ccsd(t)} + [OH3+] theory{ccsd(t)} + [SHE] theory{ccsd(t)} --> CC=C theory{ccsd(t)} + water theory{ccsd(t)}"
1905 -36.076 -28.971 -19.939 0.019 -19.920 AB + CD --> CABD "CC=C theory{ccsd(t)} + [HH] theory{ccsd(t)} --> CCC theory{ccsd(t)}"
1901 -60.004 -59.927 -68.057 -78.137 -47.594 AB --> A + B "[CH2]C(C)Cl mult{2} theory{ccsd(t)} + [SHE] theory{ccsd(t)} --> CC=C theory{ccsd(t)} + [Cl-] theory{ccsd(t)}"
1900 -60.004 -59.927 -68.057 -78.137 -47.594 AB --> A + B "[CH2]C(C)Cl mult{2} theory{ccsd(t)} + [SHE] theory{ccsd(t)} --> CC=C theory{ccsd(t)} + [Cl-] theory{ccsd(t)}"
1896 -63.046 -63.151 -71.016 -78.792 -51.207 ABCD --> BCA + D "[CH2]CCCl mult{2} theory{ccsd(t)} + [SHE] theory{ccsd(t)} --> CC=C theory{ccsd(t)} + [Cl-] theory{ccsd(t)}"
1892 -406.228 -399.225 -392.042 257.742 -35.700 A + B --> AB "C=C[CH2] mult{2} theory{ccsd(t)} + [H+] theory{ccsd(t)} + [SHE] theory{ccsd(t)} --> CC=C theory{ccsd(t)}"
1877 -60.831 -60.702 -68.664 -77.706 -47.769 AB --> A + B "[CH2]C(C)Cl mult{2} xc{pbe} + [SHE] xc{pbe} --> CC=C xc{pbe} + [Cl-] xc{pbe}"
1876 -60.831 -60.702 -68.664 -77.706 -47.769 AB --> A + B "[CH2]C(C)Cl mult{2} xc{pbe} + [SHE] xc{pbe} --> CC=C xc{pbe} + [Cl-] xc{pbe}"
1869 -67.063 -66.950 -75.120 -79.745 -56.265 ABCD --> BCA + D "[CH2]CCCl mult{2} xc{m06-2x} + [SHE] xc{m06-2x} --> CC=C xc{m06-2x} + [Cl-] xc{m06-2x}"
1868 -64.850 -64.856 -72.018 -78.650 -52.068 ABCD --> BCA + D "[CH2]CCCl mult{2} xc{pbe} + [SHE] xc{pbe} --> CC=C xc{pbe} + [Cl-] xc{pbe}"
1867 -62.558 -62.487 -70.180 -79.684 -51.264 ABCD --> BCA + D "[CH2]CCCl mult{2} xc{pbe0} + [SHE] xc{pbe0} --> CC=C xc{pbe0} + [Cl-] xc{pbe0}"
1866 -72.159 -72.264 -80.128 -78.792 -60.320 ABCD --> BCA + D "[CH2]CCCl mult{2} xc{b3lyp} + [SHE] xc{b3lyp} --> CC=C xc{b3lyp} + [Cl-] xc{b3lyp}"
1826 -63.474 -63.247 -71.605 -79.283 -52.288 AB --> A + B "[CH2]C(C)Cl mult{2} xc{m06-2x} + [SHE] xc{m06-2x} --> CC=C xc{m06-2x} + [Cl-] xc{m06-2x}"
1825 -59.426 -59.344 -67.509 -78.912 -47.821 AB --> A + B "[CH2]C(C)Cl mult{2} xc{pbe0} + [SHE] xc{pbe0} --> CC=C xc{pbe0} + [Cl-] xc{pbe0}"
1824 -68.414 -68.337 -76.466 -78.137 -56.003 AB --> A + B "[CH2]C(C)Cl mult{2} xc{b3lyp} + [SHE] xc{b3lyp} --> CC=C xc{b3lyp} + [Cl-] xc{b3lyp}"
1823 -68.414 -68.337 -76.466 -78.137 -56.003 AB --> A + B "[CH2]C(C)Cl mult{2} xc{b3lyp} + [SHE] xc{b3lyp} --> CC=C xc{b3lyp} + [Cl-] xc{b3lyp}"
1718 2.561 2.662 2.880 0.055 2.935 ABC + DE --> DBE + AC "CC=C theory{ccsd(t)} + CC theory{ccsd(t)} --> CCC theory{ccsd(t)} + C=C theory{ccsd(t)}"
1717 2.561 2.662 2.880 0.055 2.935 ABC + DE --> DBE + AC "CC=C theory{ccsd(t)} + CC theory{ccsd(t)} --> CCC theory{ccsd(t)} + C=C theory{ccsd(t)}"
1683 -46.041 -46.655 -56.218 12.519 -43.700 ABCD + E --> A + BC + DE "CCCCl xc{m06-2x} + [OH-] xc{m06-2x} --> C=CC xc{m06-2x} + O xc{m06-2x} + [Cl-] xc{m06-2x}"
985 -429.770 -422.512 -414.948 258.148 -58.201 A + B --> AB "CC=[CH] xc{m06-2x} + [H+] xc{m06-2x} + [ SHE] xc{m06-2x} --> CC=C xc{m06-2x}"
983 -258.743 -258.323 -257.944 89.336 -70.007 AB + C --> AC + B "CC=[CH] xc{m06-2x} + [OH3+] xc{m06-2x} + [ SHE] xc{m06-2x} --> CC=C xc{m06-2x} + water xc{m06-2x}"
982 -425.926 -418.630 -410.972 258.118 -54.254 A + B --> AB "C[C]=C xc{m06-2x} + [H+] xc{m06-2x} + [ SHE] xc{m06-2x} --> CC=C xc{m06-2x}"
981 -426.911 -419.703 -412.047 258.237 -55.211 A + B --> AB "C[C]=C xc{b3lyp} + [H+] xc{b3lyp} + [ SHE] xc{b3lyp} --> CC=C xc{b3lyp}"
970 -254.899 -254.442 -253.967 89.307 -66.061 AB + C --> AC + B "C[C]=C xc{m06-2x} + [OH3+] xc{m06-2x} + [ SHE] xc{m06-2x} --> CC=C xc{m06-2x} + water xc{m06-2x}"
967 -256.155 -255.749 -255.276 89.615 -67.062 AB + C --> AC + B "C[C]=C xc{b3lyp} + [OH3+] xc{b3lyp} + [ SHE] xc{b3lyp} --> CC=C xc{b3lyp} + water xc{b3lyp}"
966 -422.889 -415.796 -407.978 258.317 -51.061 A + B --> AB "C[C]=C xc{pbe} + [H+] xc{pbe} + [ SHE] xc{pbe} --> CC=C xc{pbe}"
965 -424.563 -417.361 -409.678 258.227 -52.851 A + B --> AB "C[C]=C xc{pbe0} + [H+] xc{pbe0} + [ SHE] xc{pbe0} --> CC=C xc{pbe0}"
964 -427.840 -420.838 -413.146 258.348 -56.198 A + B --> AB "CC=[CH] xc{pbe} + [H+] xc{pbe} + [ SHE] xc{pbe} --> CC=C xc{pbe}"
963 -429.119 -421.975 -414.373 258.269 -57.504 A + B --> AB "CC=[CH] xc{pbe0} + [H+] xc{pbe0} + [ SHE] xc{pbe0} --> CC=C xc{pbe0}"
961 -256.532 -256.302 -255.888 89.748 -67.541 AB + C --> AC + B "CC=[CH] xc{pbe0} + [OH3+] xc{pbe0} + [ SHE] xc{pbe0} --> CC=C xc{pbe0} + water xc{pbe0}"
956 -59.426 -59.344 -67.509 -78.912 -47.821 AB --> A + B "[CH2]C(C)Cl xc{pbe0} + [ SHE] xc{pbe0} --> CC=C xc{pbe0} + [Cl-] xc{pbe0}"
949 -252.280 -251.907 -251.280 90.056 -62.624 AB + C --> AC + B "C[C]=C xc{pbe} + [OH3+] xc{pbe} + [ SHE] xc{pbe} --> CC=C xc{pbe} + water xc{pbe}"
948 -251.976 -251.687 -251.194 89.706 -62.887 AB + C --> AC + B "C[C]=C xc{pbe0} + [OH3+] xc{pbe0} + [ SHE] xc{pbe0} --> CC=C xc{pbe0} + water xc{pbe0}"
947 -257.231 -256.949 -256.449 90.086 -67.762 AB + C --> AC + B "CC=[CH] xc{pbe} + [OH3+] xc{pbe} + [ SHE] xc{pbe} --> CC=C xc{pbe} + water xc{pbe}"
939 -431.481 -424.316 -416.738 258.308 -59.830 A + B --> AB "CC=[CH] xc{b3lyp} + [H+] xc{b3lyp} + [ SHE] xc{b3lyp} --> CC=C xc{b3lyp}"
923 -407.354 -399.909 -392.637 257.781 -36.256 A + B --> AB "C=C[CH2] xc{m06-2x} + [H+] xc{m06-2x} + [ SHE] xc{m06-2x} --> CC=C xc{m06-2x}"
922 -403.581 -396.766 -389.516 257.701 -33.215 A + B --> AB "C=C[CH2] xc{pbe} + [H+] xc{pbe} + [ SHE] xc{pbe} --> CC=C xc{pbe}"
921 -404.748 -397.656 -390.415 257.722 -34.093 A + B --> AB "C=C[CH2] xc{pbe0} + [H+] xc{pbe0} + [ SHE] xc{pbe0} --> CC=C xc{pbe0}"
920 -406.420 -399.417 -392.233 257.742 -35.892 A + B --> AB "C=C[CH2] xc{b3lyp} + [H+] xc{b3lyp} + [ SHE] xc{b3lyp} --> CC=C xc{b3lyp}"
912 -260.724 -260.361 -259.966 89.686 -71.680 AB + C --> AC + B "CC=[CH] xc{b3lyp} + [OH3+] xc{b3lyp} + [ SHE] xc{b3lyp} --> CC=C xc{b3lyp} + water xc{b3lyp}"
892 -37.724 -29.553 -20.596 0.189 -20.407 AB + CD --> CABD "CC=C xc{m06-2x} + [HH] xc{m06-2x} --> CCC xc{m06-2x}"
891 -37.082 -30.503 -21.494 0.018 -21.476 AB + CD --> CABD "CC=C xc{pbe} + [HH] xc{pbe} --> CCC xc{pbe}"
890 -40.571 -33.543 -24.539 0.196 -24.344 AB + CD --> CABD "CC=C xc{pbe0} + [HH] xc{pbe0} --> CCC xc{pbe0}"
887 -63.474 -63.247 -71.605 -79.283 -52.288 AB --> A + B "[CH2]C(C)Cl xc{m06-2x} + [ SHE] xc{m06-2x} --> CC=C xc{m06-2x} + [Cl-] xc{m06-2x}"
884 -67.063 -66.950 -75.120 -79.745 -56.265 ABCD --> BCA + D "ClCC[CH2] xc{m06-2x} + [ SHE] xc{m06-2x} --> CC=C xc{m06-2x} + [Cl-] xc{m06-2x}"
883 -64.850 -64.856 -72.018 -78.650 -52.068 ABCD --> BCA + D "ClCC[CH2] xc{pbe} + [ SHE] xc{pbe} --> CC=C xc{pbe} + [Cl-] xc{pbe}"
882 -62.558 -62.487 -70.180 -79.684 -51.264 ABCD --> BCA + D "ClCC[CH2] xc{pbe0} + [ SHE] xc{pbe0} --> CC=C xc{pbe0} + [Cl-] xc{pbe0}"
881 -72.159 -72.264 -80.128 -78.792 -60.320 ABCD --> BCA + D "ClCC[CH2] xc{b3lyp} + [ SHE] xc{b3lyp} --> CC=C xc{b3lyp} + [Cl-] xc{b3lyp}"
872 -236.328 -235.721 -235.632 88.969 -48.063 AB + C --> AC + B "C=C[CH2] xc{m06-2x} + [OH3+] xc{m06-2x} + [ SHE] xc{m06-2x} --> CC=C xc{m06-2x} + water xc{m06-2x}"
871 -232.971 -232.878 -232.818 89.440 -44.779 AB + C --> AC + B "C=C[CH2] xc{pbe} + [OH3+] xc{pbe} + [ SHE] xc{pbe} --> CC=C xc{pbe} + water xc{pbe}"
870 -232.161 -231.983 -231.930 89.201 -44.129 AB + C --> AC + B "C=C[CH2] xc{pbe0} + [OH3+] xc{pbe0} + [ SHE] xc{pbe0} --> CC=C xc{pbe0} + water xc{pbe0}"
869 -235.664 -235.462 -235.462 89.120 -47.742 AB + C --> AC + B "C=C[CH2] xc{b3lyp} + [OH3+] xc{b3lyp} + [ SHE] xc{b3lyp} --> CC=C xc{b3lyp} + water xc{b3lyp}"
843 -34.963 -27.877 -18.843 0.019 -18.825 AB + CD --> CABD "CC=C + [3H][3H] --> CCC"
666 2.561 2.662 2.880 0.055 2.935 ABC + DE --> DBE + AC "CC=C theory{ccsd(t)} + CC theory{ccsd(t)} --> CCC theory{ccsd(t)} + C=C theory{ccsd(t)}"
664 2.924 3.025 3.243 0.055 3.298 ABC + DE --> DBE + AC "CC=C theory{mp2} + CC theory{mp2} --> CCC theory{mp2} + C=C theory{mp2}"
653 2.974 2.704 2.821 0.094 2.915 ABC + DE --> DBE + AC "CC=C xc{m06-2x} + CC xc{m06-2x} --> CCC xc{m06-2x} + C=C xc{m06-2x}"
652 3.901 3.932 4.129 0.144 4.273 ABC + DE --> DBE + AC "CC=C xc{pbe} + CC xc{pbe} --> CCC xc{pbe} + C=C xc{pbe}"
651 3.691 3.727 3.899 0.101 4.000 ABC + DE --> DBE + AC "CC=C xc{pbe0} + CC xc{pbe0} --> CCC xc{pbe0} + C=C xc{pbe0}"
650 3.523 3.601 3.817 0.064 3.881 ABC + DE --> DBE + AC "CC=C xc{b3lyp} + CC xc{b3lyp} --> CCC xc{b3lyp} + C=C xc{b3lyp}"
579 -43.338 -44.068 -53.478 12.406 -41.072 ABCD + E --> A + BC + DE "CCCCl theory{ccsd(t)} + [OH-] theory{ccsd(t)} --> C=CC theory{ccsd(t)} + O theory{ccsd(t)} + [Cl-] theory{ccsd(t)}"
571 -46.040 -46.652 -56.216 12.539 -43.677 ABCD + E --> A + BC + DE "CCCCl xc{m06-2x} + [OH-] xc{m06-2x} --> C=CC xc{m06-2x} + O xc{m06-2x} + [Cl-] xc{m06-2x}"
570 -41.169 -41.851 -51.172 11.818 -39.354 ABCD + E --> A + BC + DE "CCCCl xc{pbe} + [OH-] xc{pbe} --> C=CC xc{pbe} + O xc{pbe} + [Cl-] xc{pbe}"
569 -47.272 -48.003 -57.414 12.276 -45.137 ABCD + E --> A + BC + DE "CCCCl + [OH-] --> C=CC + O + [Cl-]"
All requests to Arrows were successful.
KEYWORDs -
reaction: :reaction
chinese_room: :chinese_room
molecule: :molecule
nmr: :nmr
predict: :predict
submitesmiles: :submitesmiles
nosubmitmissingesmiles
resubmitmissingesmiles
submitmachines: :submitmachines
useallentries
nomodelcorrect
eigenvalues: :eigenvalues
frequencies: :frequencies
nwoutput: :nwoutput
xyzfile: :xyzfile
alleigs: :alleigs
allfreqs: :allfreqs
reactionenumerate:
energytype:[erxn(gas) hrxn(gas) grxn(gas) delta_solvation grxn(aq)] :energytype
energytype:[kcal/mol kj/mol ev cm-1 ry hartree au] :energytype
tablereactions:
reaction: ... :reaction
reaction: ... :reaction
...
:tablereactions
tablemethods:
method: ... :method
method: ... :method
...
:tablemethods
:reactionenumerate
rotatebonds
xyzinput:
label: :label
xyzdata:
... xyz data ...
:xyzdata
:xyzinput
submitHf: :submitHf
nmrexp: :nmrexp
findreplace: old text | new text :findreplace
listnwjobs
fetchnwjob: :fetchnwjob
pushnwjob: :pushnwjob
printcsv: :printcsv
printeig: :printeig
printfreq: :printfreq
printxyz: :printxyz
printjobinfo: :printjobinfo
printnwout: :printnwout
badids: :badids
hup_string:
database:
table:
request_table:
listallesmiles
queuecheck
This software service and its documentation were developed at the Environmental Molecular Sciences Laboratory (EMSL) at Pacific Northwest National Laboratory, a multiprogram national laboratory, operated for the U.S. Department of Energy by Battelle under Contract Number DE-AC05-76RL01830. Support for this work was provided by the Department of Energy Office of Biological and Environmental Research, and Department of Defense environmental science and technology program (SERDP). THE SOFTWARE SERVICE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE SERVICE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE SERVICE.