 3D Periodic Editor 3D Molecular And Reaction Editor Expert Editor Microsoft Quantum Editor EMSL Aerosol Workshop Editor Manual
3D Periodic Editor 3D Molecular And Reaction Editor Expert Editor Microsoft Quantum Editor EMSL Aerosol Workshop Editor Manual Results from an EMSL Arrows Calculation
| EMSL Arrows is a revolutionary approach to materials and chemical simulations that uses NWChem and chemical computational databases to make materials and chemical modeling accessible via a broad spectrum of digital communications including posts to web APIs, social networks, and traditional email. |
Molecular modeling software has previously been extremely complex, making it prohibative to all but experts in the field, yet even experts can struggle to perform calculations. This service is designed to be used by experts and non-experts alike. Experts can carry out and keep track of large numbers of complex calculations with diverse levels of theories present in their workflows. Additionally, due to a streamlined and easy-to-use input, non-experts can carry out a wide variety of molecular modeling calculations previously not accessible to them.
The id(s) for emsiles = CC(=O)Oc1ccccc1C(=O)O theory{dft} xc{pbe0} basis{6-311++G(2d,2p)} solvation_type{COSMO} ^{0} are: 78627
Use id=% instead of esmiles to print other entries.
mformula = C9H8O4
iupac = 2-acetyloxybenzoic acid
PubChem = 2244
PubChem LCSS = 2244
cas = 50-78-2
kegg = C01405 D00109
synonyms = aspirin; ACETYLSALICYLIC ACID; 50-78-2; 2-Acetoxybenzoic acid; 2-(Acetyloxy)benzoic acid; Acetylsalicylate; O-Acetylsalicylic acid; o-Acetoxybenzoic acid; Acenterine; Acetophen; Acetosal; Acylpyrin; Easprin; Ecotrin; Salicylic acid acetate; Acetosalin; Aspirdrops; Polopiryna; Salcetogen; Aceticyl; Acetonyl; Acetylin; Acidum acetylsalicylicum; Benaspir; Colfarit; Empirin; Endydol; Measurin; Rhodine; Saletin; o-Carboxyphenyl acetate; Enterosarein; Enterosarine; Acetisal; Acetylsal; Aspirine; Bialpirinia; Entericin; Enterophen; Micristin; Pharmacin; Premaspin; Salacetin; Solpyron; Temperal; Acesal; Acisal; Asagran; Asteric; Duramax; Ecolen; Extren; Globoid; Helicon; Idragin; Rhonal; Aspro; Novid; Rheumintabletten; Yasta; Solprin acid; Benzoic acid, 2-(acetyloxy)-; Acimetten; Bialpirina; Claradin; Clariprin; Delgesic; Entrophen; Globentyl; Neuronika; Acetilum acidulatum; Cemirit; Decaten; Levius; Pirseal; Solfrin; Acetilsalicilico; Adiro; Aspec; Acetosalic acid; 2-acetyloxybenzoic acid; Dolean pH 8; Triple-sal; Spira-Dine; ZORprin; Contrheuma retard; Bi-prin; XAXA; Acido acetilsalicilico; Acide acetylsalicylique; Persistin; Bayer; 2-Carboxyphenyl acetate; A.S.A. empirin; ASA; 8-hour Bayer; Endosprin; Kapsazal; Asatard; Durlaza; Bayer Plus; Acetylsalicylsaure; Rheumin tabletten; Solprin; Tasprin; Nu-seals aspirin; Salicylic acid, acetate; Acido O-acetil-benzoico; Kyselina acetylsalicylova; 2-Acetoxybenzenecarboxylic acid; Acetylsalicylsaeure; Azetylsalizylsaeure; SP 189; St. Joseph Aspirin for Adults; A.S.A.; St. Joseph; Kyselina 2-acetoxybenzoova; AC 5230; Acetylsalicyclic acid; Acetylsalicylicum acidum; Aspropharm; Cardioaspirin; Acetard; CHEBI:15365; acetyl salicylate; S-211; ECM; NSC-27223; acide 2-(acetyloxy)benzoique; Bayer Extra Strength Aspirin for Migraine Pain; Aspirin (Standard); NSC-406186; R16CO5Y76E; DTXSID5020108; 2-(acetyloxy)benzoate; o-(Acetyloxy)benzoic acid; benzoic acid, 2-acetoxy-; BAY1019036; DTXCID50108; Acetylsalicylic acid (who-ip); AXOTAL COMPONENT ASPIRIN; AZDONE COMPONENT ASPIRIN; CODOXY COMPONENT ASPIRIN; AGGRENOX COMPONENT ASPIRIN; Aspirin form II; DUOCOVER COMPONENT ASPIRIN; EXCEDRIN COMPONENT ASPIRIN; FIORINAL COMPONENT ASPIRIN; NORGESIC COMPONENT ASPIRIN; PERCODAN COMPONENT ASPIRIN; Q-GESIC COMPONENT ASPIRIN; ROXIPRIN COMPONENT ASPIRIN; VICOPRIN COMPONENT ASPIRIN; YOSPRALA COMPONENT ASPIRIN; DUOPLAVIN COMPONENT ASPIRIN; EQUAGESIC COMPONENT ASPIRIN; INVAGESIC COMPONENT ASPIRIN; LANORINAL COMPONENT ASPIRIN; MICRAININ COMPONENT ASPIRIN; ROBAXISAL COMPONENT ASPIRIN; component of Midol; ASPIRIN COMPONENT OF AXOTAL; ASPIRIN COMPONENT OF AZDONE; ASPIRIN COMPONENT OF CODOXY; NSC27223; ORPHENGESIC COMPONENT ASPIRIN; ASPIRIN COMPONENT OF AGGRENOX; ASPIRIN COMPONENT OF DUOCOVER; ASPIRIN COMPONENT OF EXCEDRIN; ASPIRIN COMPONENT OF FIORINAL; ASPIRIN COMPONENT OF NORGESIC; ASPIRIN COMPONENT OF PERCODAN; ASPIRIN COMPONENT OF Q-GESIC; ASPIRIN COMPONENT OF ROXIPRIN; ASPIRIN COMPONENT OF VICOPRIN; ASPIRIN COMPONENT OF YOSPRALA; SYNALGOS-DC COMPONENT ASPIRIN; component of Synirin; ASPIRIN COMPONENT OF DUOPLAVIN; ASPIRIN COMPONENT OF EQUAGESIC; ASPIRIN COMPONENT OF INVAGESIC; ASPIRIN COMPONENT OF LANORINAL; ASPIRIN COMPONENT OF MICRAININ; ASPIRIN COMPONENT OF ROBAXISAL; NSC406186; component of Zactirin; MEPRO-ASPIRIN COMPONENT ASPIRIN; PERCODAN-DEMI COMPONENT ASPIRIN; PRAVIGARD PAC COMPONENT ASPIRIN; SOMA COMPOUND COMPONENT ASPIRIN; ASPIRIN COMPONENT OF ORPHENGESIC; component of Coricidin; component of Persistin; component of Robaxisal; o-Acetoxybenzoate; ASPIRIN COMPONENT OF SYNALGOS-DC; DARVON COMPOUND COMPONENT ASPIRIN; INVAGESIC FORTE COMPONENT ASPIRIN; TALWIN COMPOUND COMPONENT ASPIRIN; NCGC00015067-04; Acetysal; ACIDUM ACETYLSALICYLICUM (WHO-IP); ASPIRIN COMPONENT OF MEPRO-ASPIRIN; ASPIRIN COMPONENT OF PERCODAN-DEMI; ASPIRIN COMPONENT OF PRAVIGARD PAC; ASPIRIN COMPONENT OF SOMA COMPOUND; Istopirin; Magnecyl; Medisyl; Polopirin; ORPHENGESIC FORTE COMPONENT ASPIRIN; Ronal; ASPIRIN COMPONENT OF DARVON COMPOUND; ASPIRIN COMPONENT OF INVAGESIC FORTE; ASPIRIN COMPONENT OF TALWIN COMPOUND; ASPIRIN (MART.); ASPIRIN [MART.]; Bayer Buffered; ASPIRIN COMPONENT OF ORPHENGESIC FORTE; Aspro Clear; component of Ascodeen-30; ASPIRIN (USP-RS); ASPIRIN [USP-RS]; WLN: QVR BOV1; aspirin (acetylsalicylic acid); AcetylsalicylicAcid; Aspirina 03; Acetylsalycilic acid; acetyl salicylic acid; ASPIRIN (USP MONOGRAPH); ASPIRIN [USP MONOGRAPH]; component of Darvon with A.S.A; Bayer Aspirin 8 Hour; Asaphen; Aspalon; Asprin; Bayer Children's Aspirin; Nu-seals; component of St. Joseph Cold Tablets; Aspir-Mox; Durlaza ER; Acetylsalicylsaure [German]; CAS-50-78-2; Acetoxybenzoic acid; Acetysalicylic acid; AIN; SMR000059138; Ascoden-30; CCRIS 3243; HSDB 652; Acide acetylsalicylique [French]; Acido acetilsalicilico [Italian]; Kyselina acetylsalicylova [Czech]; Acido O-acetil-benzoico [Italian]; SR-01000075668; Kyselina 2-acetoxybenzoova [Czech]; EINECS 200-064-1; NSC 27223; Aspirin [USP:BAN:JAN]; Bayer Enteric 325 mg Regular Strength; BRN 0779271; vetality; Bay E4465; UNII-R16CO5Y76E; Bayer Enteric 81 mg Adult Low Strength; Cardioaspirina; Acetyonyl; Angettes; Asacard; Ascolong; Aspirina; AspirinChewable; Bayer Enteric 500 mg Arthritis Strength; Cardiprin; Claragine; Colsprin; Encaprin; Fasprin; Medpurine; Miniasal; Salospir; Acesan; careone aspirin; Toldex; Aramark Aspirin; Aspirin Powder; Aspirin Regimen; AspirinEC; Canine Aspirin; Coated Aspirin; Enteric Aspirin; Equate Aspirin; Leader Aspirin; Rapidol Aspirin; Rexall Aspirin; Sunmark Aspirin; Topcare Aspirin; AI3-02956; Aspirin Bolus; Aspirin Nsaid; Bayer Aspirin; Rugby Aspirin; Aspi-cor; Buffered Aspirin; Chewable Aspirin; Equaline Aspirin; Geritrex Aspirin; McKesson Aspirin; AspirinNSAID; Azetylsalizylsaure; Aspirin EC; aspirinpain relief; Childrens Aspirin; Unishield Aspirin; ASA Empirin; 1oxr; 2-Acetoxybenzoate; Aspirin 5 Grain; Care One Aspirin; Bufferin Arthritis; ULINE Aspirin; CAREALL Aspirin; MooreBrand Aspirin; Aspica (Aspirin); Aspirin 81mg; Dg Health Aspirin; Medi-first Aspirin; Basic Care Aspirin; caring mill aspirin; Dr Pausins Aspirin; Good Sense Aspirin; rugby adult aspirin; Aspirin 325mg; Aspirin 81; Aspirin 81 mg; Aspirin,(S); Up and Up Aspirin; Aspalon (JAN); Durlaza (TN); Easprin (TN); Health Mart Aspirin; MBR Aspirin Powder; Pain Relief Aspirin; Solves-aspirinCherry; Aspirin 325 mg; Aspirin Pain reliver; MFCD00002430; Tri-buffered Aspirin; acetyl-salicylic acid; Aspirin 325; Aspirin 50 CT; AspirinEnteric Coated; ASPRISOL; Crane Safety Aspirin; equate aspirinchewable; Health Sense Ecpirin; Henry Schein Aspirin; sunmark adult aspirin; Travel Savvy Aspirin; VAZALORE; ASPIRINLow Strength; acetyl salicyclic acid; Adult Aspirin Regimen; Aspirin Bolus-240; AspirinDelayed Release; Bayer Aspirin Regimen; Bayer Genuine Aspirin; Direct Safety Aspirin; Moore Medical Aspirin; o-(Acetyloxy)benzoate; Rapid Comfort Aspirin; Safety Coated Aspirin; signature care aspirin; Aspirinregular strength; ASSURED ASPIRIN; Percodan (Salt/Mix); Adult Chewable Aspirin; Biovanta Double Action; Enteric Coated Aspirin; Extra Strength Aspirin; ADVANCED ASPIRIN; Ascriptin (Salt/Mix); Micrainin (Salt/Mix); VALUMEDS ASPIRIN; 2-acetoxy benzoic acid; Aspirin tablet 325mg; Chewable Aspirin 81mg; Dye-Free Aspirin 81; Family Wellness Aspirin; Physicians Care Aspirin; RHODINE NC RP; Signature Care Aspririn; Spectrum_001245; 2-Acetylsalicyclic acid; First Aid Only Aspirin; Acide acetyl salicylique; ASPIRIN [VANDF]; Medique at Home Aspirin; ASPIRIN [HSDB]; Aspirin Regular Strength; Ecotrin Regular Strength; Medi-First Plus Aspirin; Medique Products Aspirin; Regular Strength Aspirin; Salicylic acid, acetyl-; ASPIRIN [JAN]; Bayer PlusExtra Strength; Circle K Aspirin 325; Pharbest Aspirin 325mg; ASPIRIN [MI]; CHEMBL25; health mart adult aspirin; Spectrum2_001899; Spectrum3_001295; Spectrum4_000099; Spectrum5_000740; Aspirin (JP17/USP); Lopac-A-5376; Salycylacetylsalicylic acid; ASPIRIN480; BAYER 500 mg; Chronic Pain/Fever Relief; CARDIASPIRIN PROTECT; NobleAid PAIN RELIEVER; Berkley and Jensen Aspirin; Epitope ID:114151; Percodan Demi (Salt/Mix); Soma Compound (Salt/Mix); EC 200-064-1; Acetylsalicylic acid, 99%; ASPIRINEXTRA STRENGTH; cid_2244; Pravigard PAC (Salt/Mix); SCHEMBL1353; Up and Up Chewable Aspirin; ASPIRIN BOLUS-480; AspirinEnteric Safety Coated; AspirinEnteric Safety-Coated; Regular Strength Aspirin EC; 2-(Acetyloxy)-benzoic acid; Bay-e-4465; Plus PharmaNSAID 325 mg; 365 Everyday Value Aspirin; Alka-Seltzer Original Flavor; Aspirin 81mg Enteric coated; AspirinLow Strength, Enteric; Critical Care Aspirin To Go; Lopac0_000038; ASPIRIN 325 MG EC; KBioGR_000398; KBioGR_002271; KBioSS_001725; KBioSS_002272; 4-10-00-00138 (Beilstein Handbook Reference); MLS001055329; MLS001066332; MLS001336045; MLS001336046; ASPIRIN [ORANGE BOOK]; BAYER Aspirin Extra Strength; BIDD:GT0118; DivK1c_000555; Lil Drug Store Aspirin 325; SPECTRUM1500130; Aspirin 81 mg Enteric Coated; Bayer Aspirin Regimen Chewable; Bayer Chewable-Aspirin Regimen; Good Neighbor Pharmacy Aspirin; Rapidol Aspirin Display 2x25; SPBio_001838; Acetylsalicylic acid, >=99%; Buffered AspirinFor Small Dogs; GTPL4139; O-Acetylsalicylic acid; Aspirin; MBR Aspirin Bolus 240 Grains; BDBM22360; HMS501L17; KBio1_000555; KBio2_001725; KBio2_002271; KBio2_004293; KBio2_004839; KBio2_006861; KBio2_007407; KBio3_002149; KBio3_002751; Empirin with Codeine (Salt/Mix); Pharbest Regular Strength Aspirin; Value PharmaAspirin Pain Reliever; Acetylsalicylic acid, >=99.0%; cMAP_000006; component of Zactirin (Salt/Mix); First Aid Direct Chewable Aspirin; NINDS_000555; ASPIRIN Analgesic and Antipyretic; HMS1920E13; HMS2090G03; HMS2091K13; HMS2233L18; HMS3260G17; HMS3372N15; HMS3656N14; HMS3715P19; HMS3866L03; HMS3885G03; Pharmakon1600-01500130; ACETYLSALICYLIC ACID [INCI]; Bayer Aspirin Regimenenteric coated; BCP21790; STR01551; ACETYLSALICYLIC ACID; ASPIRIN; BAYER AspirinExtra Strength Caplets; Tox21_110076; Tox21_202117; Tox21_300146; Tox21_500038; Buffered aspirin, effervescent tablet; CCG-39490; HY-14654R; NSC755899; s3017; ACETYLSALICYLIC ACID [WHO-DD]; Bufferin Regular Strength Pain Relief; enteric coated aspirinRegular Strength; AKOS000118884; Aspirin Enteric Coated Tablets 81 mg; component of Ascodeen-30 (Salt/Mix); Tox21_110076_1; ACETYLSALICYLIC ACID [EMA EPAR]; ACETYLSALICYLICUM ACIDUM [HPUS]; Aspirin 81 mg Delayed Release Tablets; CS-2001; DB00945; LP00038; NSC-755899; PL-2200; SDCCGSBI-0050027.P005; Value PharmaPain RelieverExtra Strength; AspirinEnteric Coated, Regular Strength; BAY-1019036; IDI1_000555; Regular Strength Enteric Coated Aspirin; ACETYLSALICYLIC ACID [GREEN BOOK]; Acetylsalicylic acid, analytical standard; Aspirin Delayed Release Tablets, 81 mg; Buffered AspirinFor Medium to Large Dogs; Coraspirin 81 mg Enteric Coasted Tablet; NCGC00015067-01; NCGC00015067-02; NCGC00015067-03; NCGC00015067-05; NCGC00015067-06; NCGC00015067-07; NCGC00015067-08; NCGC00015067-09; NCGC00015067-10; NCGC00015067-11; NCGC00015067-12; NCGC00015067-13; NCGC00015067-14; NCGC00015067-24; NCGC00015067-26; NCGC00090977-01; NCGC00090977-02; NCGC00090977-03; NCGC00090977-04; NCGC00090977-05; NCGC00090977-06; NCGC00090977-07; NCGC00254034-01; NCGC00259666-01; NCGC00260723-01; Aspirin, meets USP testing specifications; HY-14654; NCI60_002222; ACETYLSALICYLIC ACID [EP MONOGRAPH]; Aspirin 81mg Enteric coatedDelayed Release; Cardioaspirin 81 mg Enteric Coated Tablet; SBI-0050027.P004; UNM-0000306102; component of Darvon with A.S.A (Salt/Mix); CS-0694916; EU-0100038; FT-0655181; FT-0661360; SW199665-2; CARISOPRODOL COMPOUND COMPONENT ASPIRIN; EN300-19606; A 5376; Acetylsalicylic Acid 1.0 mg/ml in Acetonitrile; C01405; D00109; E80792; Q18216; Safety Coated Aspirin 325 mg Regular Strength; AB00051918-08; AB00051918_09; AB00051918_10; ASPIRIN COMPONENT OF CARISOPRODOL COMPOUND; Arthritis Pain Formula Maximum Strength (Salt/Mix); Health Mart Regular Strength Enteric Coated Aspirin; Natural Aspirin plus Tart Cherry Dietary Supplement; SR-01000075668-1; SR-01000075668-4; SR-01000075668-6; Acetylsalicylic acid, Vetec(TM) reagent grade, >=99%; Aspirin 81mg Enteric coatedLow Strength Aspirin Regimen; Aspirin, British Pharmacopoeia (BP) Reference Standard; CLOPIDOGREL/ACETYLSALICYLIC ACID COMPONENT ASPIRIN; F2191-0068; Natural Aspirin plus Immune Supporting Dietary Supplement; Natural Aspirin plus Lemon and Honey Dietary Supplement; Z104474430; ASPIRIN COMPONENT OF CLOPIDOGREL/ACETYLSALICYLIC ACID; Aspirin, United States Pharmacopeia (USP) Reference Standard; D41527A7-A9EB-472D-A7FC-312821130549; Acetylsalicylic acid, European Pharmacopoeia (EP) Reference Standard; Acetylsalicylic acid, BioReagent, plant cell culture tested, >=99.0%; Acetylsalicylic acid for peak identification, European Pharmacopoeia (EP) Reference Standard; InChI=1/C9H8O4/c1-6(10)13-8-5-3-2-4-7(8)9(11)12/h2-5H,1H3,(H,11,12; 11126-35-5; Aspirin (Acetyl Salicylic Acid), Pharmaceutical Secondary Standard; Certified Reference Material
Search Links to Other Online Resources (may not be available):
- Google Structure Search
- EPA CompTox Database
- Chemical Entities of Biological Interest (ChEBI)
- NIH ChemIDplus - A TOXNET DATABASE
- The Human Metabolome Database (HMDB)
- OECD eChemPortal
- Google Scholar
+==================================================+
|| Molecular Calculation ||
+==================================================+
Id = 78627
NWOutput = Link to NWChem Output (download)
Datafiles:
lumo-restricted.cube-847896-2023-11-20-2:37:2 (download)
homo-restricted.cube-847896-2023-11-20-2:37:2 (download)
cosmo.xyz-847896-2023-11-20-2:37:2 (download)
mo_orbital_nwchemarrows.out-805770-2024-2-18-5:37:1 (download)
image_resset: api/image_reset/78627
Calculation performed by Eric Bylaska - arrow6.emsl.pnl.gov
Numbers of cpus used for calculation = 32
Calculation walltime = 51451.400000 seconds (0 days 14 hours 17 minutes 31 seconds)
+----------------+
| Energetic Data |
+----------------+
Id = 78627
iupac = 2-acetyloxybenzoic acid
mformula = C9H8O4
inchi = InChI=1S/C9H8O4/c1-6(10)13-8-5-3-2-4-7(8)9(11)12/h2-5H,1H3,(H,11,12)
inchikey = BSYNRYMUTXBXSQ-UHFFFAOYSA-N
esmiles = CC(=O)Oc1ccccc1C(=O)O theory{dft} xc{pbe0} basis{6-311++G(2d,2p)} solvation_type{COSMO} ^{0}
calculation_type = ovc
theory = dft
xc = pbe0
basis = 6-311++G(2d,2p)
charge,mult = 0 1
energy = -648.178809 Hartrees
enthalpy correct.= 0.169798 Hartrees
entropy = 103.160 cal/mol-K
solvation energy = -14.300 kcal/mol solvation_type = COSMO
Sitkoff cavity dispersion = 2.650 kcal/mol
Honig cavity dispersion = 8.948 kcal/mol
ASA solvent accesible surface area = 357.939 Angstrom2
ASA solvent accesible volume = 335.823 Angstrom3
+-----------------+
| Structural Data |
+-----------------+
JSmol: an open-source HTML5 viewer for chemical structures in 3D
Number of Atoms = 21
Units are Angstrom for bonds and degrees for angles
Type I J K L M Value
----------- ----- ----- ----- ----- ----- ----------
1 Stretch C1 C2 1.49314
2 Stretch C1 H14 1.08995
3 Stretch C1 H15 1.09190
4 Stretch C1 H16 1.08564
5 Stretch C2 O3 1.18994
6 Stretch C2 O4 1.38177
7 Stretch O4 C5 1.39399
8 Stretch C5 C6 1.38345
9 Stretch C5 C10 1.39473
10 Stretch C6 C7 1.38398
11 Stretch C6 H17 1.08059
12 Stretch C7 C8 1.38859
13 Stretch C7 H18 1.08265
14 Stretch C8 C9 1.38120
15 Stretch C8 H19 1.08217
16 Stretch C9 C10 1.39453
17 Stretch C9 H20 1.08190
18 Stretch C10 C11 1.50191
19 Stretch C11 O12 1.19915
20 Stretch C11 O13 1.33895
21 Stretch O13 H21 0.96736
22 Bend C2 C1 H14 110.43546
23 Bend C2 C1 H15 109.55782
24 Bend C2 C1 H16 109.23244
25 Bend H14 C1 H15 107.61714
26 Bend H14 C1 H16 110.53464
27 Bend H15 C1 H16 109.43840
28 Bend C1 C2 O3 127.05327
29 Bend C1 C2 O4 109.79646
30 Bend O3 C2 O4 123.14912
31 Bend C2 O4 C5 119.15134
32 Bend O4 C5 C6 119.53183
33 Bend O4 C5 C10 118.81321
34 Bend C6 C5 C10 121.57899
35 Bend C5 C6 C7 119.45151
36 Bend C5 C6 H17 119.59397
37 Bend C7 C6 H17 120.95181
38 Bend C6 C7 C8 120.19104
39 Bend C6 C7 H18 119.43700
40 Bend C8 C7 H18 120.37194
41 Bend C7 C8 C9 119.70247
42 Bend C7 C8 H19 120.28332
43 Bend C9 C8 H19 120.01394
44 Bend C8 C9 C10 121.33033
45 Bend C8 C9 H20 121.55188
46 Bend C10 C9 H20 117.11779
47 Bend C5 C10 C9 117.74260
48 Bend C5 C10 C11 126.12754
49 Bend C9 C10 C11 116.12868
50 Bend C10 C11 O12 121.78195
51 Bend C10 C11 O13 117.68994
52 Bend O12 C11 O13 120.52017
53 Bend C11 O13 H21 109.27782
54 Dihedral C1 C2 O4 C5 177.71338
55 Dihedral C2 O4 C5 C6 59.89465
56 Dihedral C2 O4 C5 C10 -123.22421
57 Dihedral O3 C2 C1 H14 -128.85424
58 Dihedral O3 C2 C1 H15 112.77824
59 Dihedral O3 C2 C1 H16 -7.08845
60 Dihedral O3 C2 O4 C5 -1.92200
61 Dihedral O4 C2 C1 H14 51.52828
62 Dihedral O4 C2 C1 H15 -66.83924
63 Dihedral O4 C2 C1 H16 173.29408
64 Dihedral O4 C5 C6 C7 177.30639
65 Dihedral O4 C5 C6 H17 -3.28600
66 Dihedral O4 C5 C10 C9 -177.48238
67 Dihedral O4 C5 C10 C11 2.10631
68 Dihedral C5 C6 C7 C8 -0.05841
69 Dihedral C5 C6 C7 H18 179.89695
70 Dihedral C5 C10 C9 C8 0.38119
71 Dihedral C5 C10 C9 H20 -179.63970
72 Dihedral C5 C10 C11 O12 170.78775
73 Dihedral C5 C10 C11 O13 -10.23317
74 Dihedral C6 C5 C10 C9 -0.66781
75 Dihedral C6 C5 C10 C11 178.92089
76 Dihedral C6 C7 C8 C9 -0.21974
77 Dihedral C6 C7 C8 H19 179.96921
78 Dihedral C7 C6 C5 C10 0.51422
79 Dihedral C7 C8 C9 C10 0.05236
80 Dihedral C7 C8 C9 H20 -179.92583
81 Dihedral C8 C7 C6 H17 -179.45777
82 Dihedral C8 C9 C10 C11 -179.24879
83 Dihedral C9 C8 C7 H18 179.82533
84 Dihedral C9 C10 C11 O12 -9.61771
85 Dihedral C9 C10 C11 O13 169.36138
86 Dihedral C10 C5 C6 H17 179.92184
87 Dihedral C10 C9 C8 H19 179.86393
88 Dihedral C10 C11 O13 H21 1.26859
89 Dihedral C11 C10 C9 H20 0.73033
90 Dihedral O12 C11 O13 H21 -179.73882
91 Dihedral H17 C6 C7 H18 0.49759
92 Dihedral H18 C7 C8 H19 0.01427
93 Dihedral H19 C8 C9 H20 -0.11426
+-----------------+
| Eigenvalue Data |
+-----------------+
Id = 78627
iupac = 2-acetyloxybenzoic acid
mformula = C9H8O4
InChI = InChI=1S/C9H8O4/c1-6(10)13-8-5-3-2-4-7(8)9(11)12/h2-5H,1H3,(H,11,12)
smiles = CC(=O)Oc1ccccc1C(=O)O
esmiles = CC(=O)Oc1ccccc1C(=O)O theory{dft} xc{pbe0} basis{6-311++G(2d,2p)} solvation_type{COSMO} ^{0}
theory = dft
xc = pbe0
basis = 6-311++G(2d,2p)
charge = 0
mult = 1
solvation_type = COSMO
twirl webpage = TwirlMol Link
image webpage = GIF Image Link
Eigevalue Spectra
---- ---- 67.36 eV
---- ----
----------
---- ----
---- ----
--- -- ---
--- -- ---
--- -- ---
---- ----
--- -- ---
--- -- ---
-- -- -- -
- - - - --
6 - - - -
9 - - - -
8 - - - -
8 - - - -
8 - - - -
10 - - - -
9 - - - -
6 - - - -
6 - - - -
- - - - --
7 - - - -
12 - - - -
9 - - - -
11 - - - -
8 - - - -
8 - - - -
14 - - - -
13 - - - -
17 - - - -
13 - - - -
9 - - - -
---- ---- LUMO= -1.76 eV
HOMO= -7.65 eV ++++ ++++
6 + + + +
6 + + + +
7 + + + +
++++++++++
++ ++ ++ +
+++ ++ +++
++++++++++
++++ ++++
-32.09 eV ++++ ++++
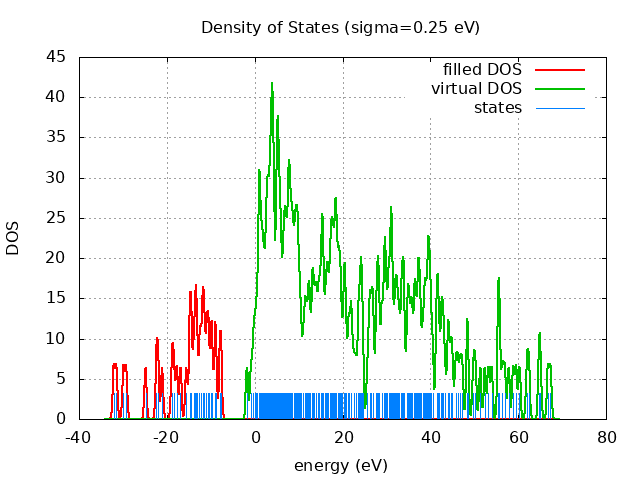
spin eig occ ---------------------------- restricted -32.09 2.00 restricted -31.50 2.00 restricted -29.89 2.00 restricted -29.27 2.00 restricted -24.86 2.00 restricted -22.36 2.00 restricted -22.02 2.00 restricted -21.08 2.00 restricted -18.83 2.00 restricted -18.44 2.00 restricted -17.74 2.00 restricted -16.89 2.00 restricted -15.58 2.00 restricted -14.82 2.00 restricted -14.60 2.00 restricted -14.43 2.00 restricted -13.89 2.00 restricted -13.52 2.00 restricted -13.37 2.00 restricted -13.14 2.00 restricted -12.56 2.00 restricted -12.27 2.00 restricted -11.83 2.00 restricted -11.72 2.00 restricted -11.45 2.00 restricted -11.01 2.00 restricted -10.70 2.00 restricted -10.46 2.00 restricted -9.90 2.00 restricted -9.72 2.00 restricted -9.02 2.00 restricted -8.84 2.00 restricted -7.92 2.00 restricted -7.65 2.00 restricted -1.76 0.00 restricted -0.83 0.00 restricted -0.30 0.00 restricted -0.03 0.00 restricted 0.32 0.00 restricted 0.52 0.00 restricted 0.91 0.00 restricted 0.97 0.00 restricted 1.03 0.00 restricted 1.10 0.00 restricted 1.29 0.00 restricted 1.48 0.00 restricted 1.67 0.00 restricted 1.77 0.00 restricted 1.96 0.00 restricted 2.22 0.00 restricted 2.28 0.00 restricted 2.64 0.00 restricted 2.68 0.00 restricted 2.84 0.00 restricted 2.94 0.00 restricted 3.08 0.00 restricted 3.24 0.00 restricted 3.34 0.00 restricted 3.51 0.00 restricted 3.62 0.00 restricted 3.78 0.00 restricted 3.78 0.00 restricted 3.92 0.00 restricted 4.04 0.00 restricted 4.08 0.00 restricted 4.16 0.00 restricted 4.25 0.00 restricted 4.41 0.00 restricted 4.54 0.00 restricted 4.90 0.00 restricted 4.97 0.00 restricted 5.13 0.00 restricted 5.19 0.00 restricted 5.24 0.00 restricted 5.36 0.00 restricted 5.48 0.00 restricted 5.57 0.00 restricted 5.81 0.00 restricted 5.86 0.00 restricted 6.00 0.00 restricted 6.31 0.00 restricted 6.47 0.00 restricted 6.72 0.00 restricted 6.75 0.00 restricted 6.95 0.00 restricted 7.13 0.00 restricted 7.20 0.00 restricted 7.45 0.00 restricted 7.61 0.00 restricted 7.71 0.00 restricted 7.86 0.00 restricted 7.93 0.00 restricted 8.03 0.00 restricted 8.19 0.00 restricted 8.44 0.00 restricted 8.54 0.00 restricted 8.56 0.00 restricted 8.89 0.00 restricted 8.95 0.00 restricted 9.19 0.00 restricted 9.32 0.00 restricted 9.46 0.00 restricted 9.63 0.00 restricted 9.77 0.00 restricted 9.88 0.00 restricted 10.12 0.00 restricted 10.31 0.00 restricted 10.59 0.00 restricted 11.03 0.00 restricted 11.35 0.00 restricted 11.59 0.00 restricted 11.85 0.00 restricted 12.15 0.00 restricted 12.45 0.00 restricted 12.49 0.00 restricted 13.02 0.00 restricted 13.12 0.00 restricted 13.34 0.00 restricted 13.60 0.00 restricted 13.86 0.00 restricted 14.05 0.00 restricted 14.32 0.00 restricted 14.62 0.00 restricted 14.75 0.00 restricted 15.06 0.00 restricted 15.23 0.00 restricted 15.41 0.00 restricted 15.53 0.00 restricted 15.55 0.00 restricted 15.99 0.00 restricted 16.18 0.00 restricted 16.45 0.00 restricted 16.60 0.00 restricted 16.79 0.00 restricted 17.10 0.00 restricted 17.25 0.00 restricted 17.46 0.00 restricted 17.52 0.00 restricted 17.77 0.00 restricted 17.84 0.00 restricted 18.06 0.00 restricted 18.27 0.00 restricted 18.41 0.00 restricted 18.52 0.00 restricted 18.58 0.00 restricted 18.87 0.00 restricted 19.03 0.00 restricted 19.25 0.00 restricted 19.36 0.00 restricted 19.63 0.00 restricted 19.80 0.00 restricted 20.32 0.00 restricted 20.35 0.00 restricted 20.59 0.00 restricted 20.81 0.00 restricted 21.32 0.00 restricted 21.53 0.00 restricted 21.98 0.00 restricted 22.02 0.00 restricted 22.55 0.00 restricted 23.03 0.00 restricted 23.59 0.00 restricted 23.74 0.00 restricted 24.10 0.00 restricted 24.20 0.00 restricted 24.39 0.00 restricted 24.62 0.00 restricted 25.69 0.00 restricted 26.18 0.00 restricted 26.24 0.00 restricted 26.59 0.00 restricted 26.86 0.00 restricted 26.99 0.00 restricted 27.61 0.00 restricted 27.84 0.00 restricted 28.05 0.00 restricted 28.19 0.00 restricted 28.38 0.00 restricted 28.90 0.00 restricted 28.98 0.00 restricted 29.44 0.00 restricted 29.54 0.00 restricted 29.70 0.00 restricted 29.84 0.00 restricted 30.15 0.00 restricted 30.49 0.00 restricted 30.51 0.00 restricted 30.84 0.00 restricted 31.06 0.00 restricted 31.13 0.00 restricted 31.17 0.00 restricted 31.41 0.00 restricted 31.80 0.00 restricted 32.00 0.00 restricted 32.27 0.00 restricted 32.39 0.00 restricted 32.74 0.00 restricted 32.95 0.00 restricted 33.36 0.00 restricted 33.52 0.00 restricted 33.76 0.00 restricted 33.94 0.00 restricted 34.07 0.00 restricted 34.69 0.00 restricted 34.88 0.00 restricted 35.10 0.00 restricted 35.37 0.00 restricted 35.71 0.00 restricted 35.86 0.00 restricted 36.28 0.00 restricted 36.53 0.00 restricted 36.62 0.00 restricted 37.04 0.00 restricted 37.21 0.00 restricted 37.40 0.00 restricted 37.56 0.00 restricted 37.85 0.00 restricted 38.26 0.00 restricted 38.53 0.00 restricted 38.86 0.00 restricted 38.91 0.00 restricted 39.30 0.00 restricted 39.45 0.00 restricted 39.59 0.00 restricted 39.77 0.00 restricted 39.95 0.00 restricted 40.23 0.00 restricted 40.56 0.00 restricted 41.42 0.00 restricted 41.48 0.00 restricted 41.76 0.00 restricted 41.98 0.00 restricted 42.45 0.00 restricted 42.66 0.00 restricted 42.91 0.00 restricted 43.28 0.00 restricted 43.97 0.00 restricted 44.16 0.00 restricted 44.65 0.00 restricted 45.00 0.00 restricted 45.76 0.00 restricted 46.23 0.00 restricted 46.78 0.00 restricted 47.26 0.00 restricted 48.38 0.00 restricted 48.50 0.00 restricted 49.81 0.00 restricted 50.25 0.00 restricted 51.36 0.00 restricted 52.41 0.00 restricted 53.24 0.00 restricted 53.95 0.00 restricted 55.44 0.00 restricted 55.59 0.00 restricted 55.70 0.00 restricted 56.38 0.00 restricted 56.94 0.00 restricted 57.83 0.00 restricted 58.86 0.00 restricted 59.50 0.00 restricted 60.29 0.00 restricted 62.00 0.00 restricted 62.44 0.00 restricted 64.75 0.00 restricted 65.05 0.00 restricted 66.77 0.00 restricted 67.36 0.00
+----------------------------------------+ | Vibrational Density of States Analysis | +----------------------------------------+ Total number of frequencies = 63 Total number of negative frequencies = 1 - w_negative = -58.7 cm-1 Number of lowest frequencies = 11 (frequency threshold = 500 ) Exact dos norm = 57.000 Generating vibrational DOS Generating model vibrational DOS to have a proper norm 10.00 56.00 11.00 57.00 50.00 55.79 10.79 57.00 100.00 55.16 10.16 57.00 Generating IR Spectra Writing vibrational density of states (DOS) to vdos.dat Writing model vibrational density of states (DOS_FIXED) to vdos-model.dat Writing IR spectra to irdos.dat Temperature= 298.15 zero-point correction to energy = 99.210 kcal/mol ( 0.158102) vibrational contribution to enthalpy correction = 104.181 kcal/mol ( 0.166023) vibrational contribution to Entropy = 30.582 cal/mol-k hindered rotor enthalpy correction = 0.000 kcal/mol ( 0.000000) hindered rotor entropy correction = 0.000 cal/mol-k
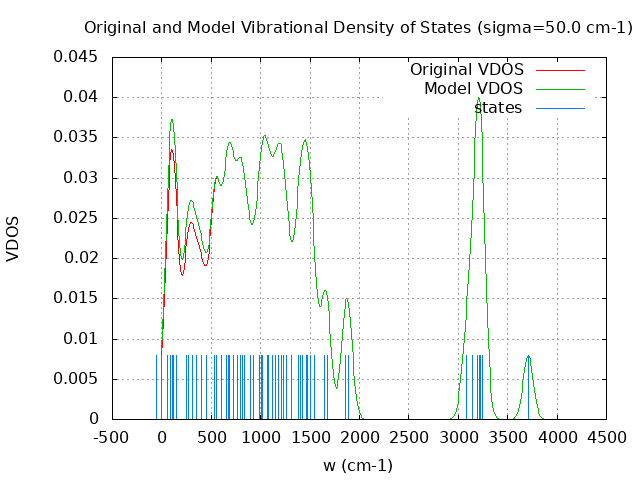
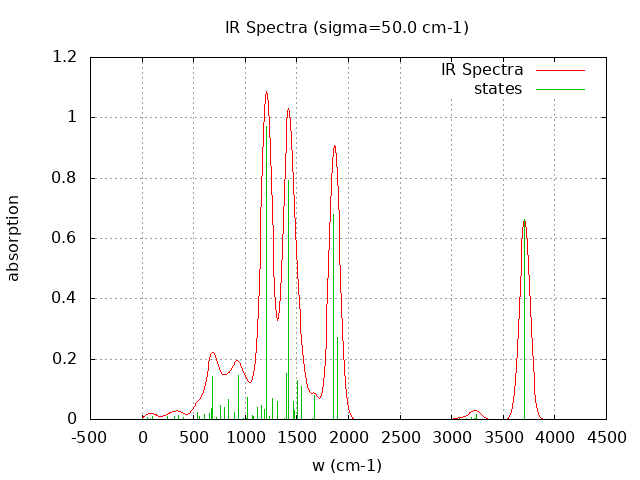
DOS sigma = 10.000000
- vibrational DOS enthalpy correction = 0.166026 kcal/mol ( 104.183 kcal/mol)
- model vibrational DOS enthalpy correction = 0.167091 kcal/mol ( 104.851 kcal/mol)
- vibrational DOS Entropy = 0.000049 ( 30.662 cal/mol-k)
- model vibrational DOS Entropy = 0.000053 ( 32.961 cal/mol-k)
- original gas Energy = -648.178809 (-406738.341 kcal/mol)
- original gas Enthalpy = -648.009011 (-406631.791 kcal/mol, delta= 0.000)
- unajusted DOS gas Enthalpy = -648.009009 (-406631.789 kcal/mol, delta= 0.002)
- model DOS gas Enthalpy = -648.007943 (-406631.120 kcal/mol, delta= 0.670)
- original gas Entropy = 0.000164 ( 103.160 cal/mol-k,delta= 0.000)
- unajusted DOS gas Entropy = 0.000165 ( 103.240 cal/mol-k,delta= 0.080)
- model DOS gas Entropy = 0.000168 ( 105.538 cal/mol-k,delta= 2.378)
- original gas Free Energy = -648.058026 (-406662.548 kcal/mol, delta= 0.000)
- unadjusted DOS gas Free Energy = -648.058061 (-406662.570 kcal/mol, delta= -0.022)
- model DOS gas Free Energy = -648.058088 (-406662.587 kcal/mol, delta= -0.039)
- original sol Free Energy = -648.080815 (-406676.848 kcal/mol)
- unadjusted DOS sol Free Energy = -648.080850 (-406676.870 kcal/mol)
- model DOS sol Free Energy = -648.080877 (-406676.887 kcal/mol)
DOS sigma = 50.000000
- vibrational DOS enthalpy correction = 0.165931 kcal/mol ( 104.124 kcal/mol)
- model vibrational DOS enthalpy correction = 0.167229 kcal/mol ( 104.938 kcal/mol)
- vibrational DOS Entropy = 0.000051 ( 31.742 cal/mol-k)
- model vibrational DOS Entropy = 0.000055 ( 34.682 cal/mol-k)
- original gas Energy = -648.178809 (-406738.341 kcal/mol)
- original gas Enthalpy = -648.009011 (-406631.791 kcal/mol, delta= 0.000)
- unajusted DOS gas Enthalpy = -648.009103 (-406631.848 kcal/mol, delta= -0.058)
- model DOS gas Enthalpy = -648.007805 (-406631.034 kcal/mol, delta= 0.757)
- original gas Entropy = 0.000164 ( 103.160 cal/mol-k,delta= 0.000)
- unajusted DOS gas Entropy = 0.000166 ( 104.320 cal/mol-k,delta= 1.160)
- model DOS gas Entropy = 0.000171 ( 107.260 cal/mol-k,delta= 4.100)
- original gas Free Energy = -648.058026 (-406662.548 kcal/mol, delta= 0.000)
- unadjusted DOS gas Free Energy = -648.058669 (-406662.951 kcal/mol, delta= -0.403)
- model DOS gas Free Energy = -648.058768 (-406663.013 kcal/mol, delta= -0.466)
- original sol Free Energy = -648.080815 (-406676.848 kcal/mol)
- unadjusted DOS sol Free Energy = -648.081458 (-406677.252 kcal/mol)
- model DOS sol Free Energy = -648.081557 (-406677.314 kcal/mol)
DOS sigma = 100.000000
- vibrational DOS enthalpy correction = 0.165564 kcal/mol ( 103.893 kcal/mol)
- model vibrational DOS enthalpy correction = 0.167582 kcal/mol ( 105.160 kcal/mol)
- vibrational DOS Entropy = 0.000048 ( 29.952 cal/mol-k)
- model vibrational DOS Entropy = 0.000055 ( 34.332 cal/mol-k)
- original gas Energy = -648.178809 (-406738.341 kcal/mol)
- original gas Enthalpy = -648.009011 (-406631.791 kcal/mol, delta= 0.000)
- unajusted DOS gas Enthalpy = -648.009471 (-406632.079 kcal/mol, delta= -0.288)
- model DOS gas Enthalpy = -648.007452 (-406630.812 kcal/mol, delta= 0.978)
- original gas Entropy = 0.000164 ( 103.160 cal/mol-k,delta= 0.000)
- unajusted DOS gas Entropy = 0.000163 ( 102.530 cal/mol-k,delta= -0.630)
- model DOS gas Entropy = 0.000170 ( 106.909 cal/mol-k,delta= 3.749)
- original gas Free Energy = -648.058026 (-406662.548 kcal/mol, delta= 0.000)
- unadjusted DOS gas Free Energy = -648.058186 (-406662.648 kcal/mol, delta= -0.100)
- model DOS gas Free Energy = -648.058248 (-406662.687 kcal/mol, delta= -0.139)
- original sol Free Energy = -648.080815 (-406676.848 kcal/mol)
- unadjusted DOS sol Free Energy = -648.080975 (-406676.949 kcal/mol)
- model DOS sol Free Energy = -648.081037 (-406676.988 kcal/mol)
Normal Mode Frequency (cm-1) IR Intensity (arbitrary)
----------- ---------------- ------------------------
1 -58.720 0.078
2 0.000 0.149
3 0.000 0.107
4 0.000 0.038
5 0.000 0.567
6 0.000 0.551
7 0.000 0.495
8 57.120 0.862
9 82.450 0.460
10 102.420 1.285
11 120.340 0.046
12 147.470 0.172
13 245.480 1.220
14 267.550 0.201
15 309.880 1.352
16 353.220 1.716
17 399.580 0.975
18 449.660 0.193
19 532.400 2.000
20 534.210 2.713
21 553.640 1.316
22 604.540 2.003
23 655.360 2.336
24 676.600 4.632
25 682.180 18.010
26 723.560 0.745
27 762.300 5.997
28 797.010 5.093
29 816.220 0.090
30 839.780 8.514
31 893.050 2.924
32 929.360 18.143
33 986.870 0.440
34 1010.500 0.567
35 1021.470 9.287
36 1065.620 1.596
37 1079.100 1.060
38 1119.960 4.903
39 1154.350 5.895
40 1182.690 4.165
41 1207.520 121.804
42 1231.840 1.049
43 1263.320 8.800
44 1314.620 7.441
45 1380.130 0.363
46 1399.010 19.129
47 1419.340 99.323
48 1466.740 7.534
49 1474.740 3.872
50 1508.090 16.049
51 1541.320 13.879
52 1646.740 0.260
53 1672.860 10.142
54 1855.750 85.061
55 1889.730 34.264
56 3078.620 0.478
57 3140.520 0.261
58 3190.500 0.805
59 3196.280 0.259
60 3218.190 0.324
61 3226.960 0.466
62 3244.690 2.251
63 3708.310 83.291
No Hindered Rotor Data
+-------------------------------------+
| Reactions Contained in the Database |
+-------------------------------------+
Reactions Containing INCHIKEY = BSYNRYMUTXBXSQ-UHFFFAOYSA-N
Reactionid Erxn(gas) Hrxn(gas) Grxn(gas) delta_Solv Grxn(aq) ReactionType Reaction
20799 -2.564 -1.886 -2.662 -4.718 -7.380 AB + CD --> AD + BC "CC(=O)Oc1ccccc1C(=O)O + O --> OC(=O)c1ccccc1O + CC(=O)O"
20798 -2.564 -1.886 -2.662 -4.718 -7.380 AB + CD --> AD + BC "CC(=O)Oc1ccccc1C(=O)O + O --> OC(=O)c1ccccc1O + CC(=O)O"
20797 -2.564 -1.886 -2.662 -4.718 -7.380 AB + CD --> AD + BC "CC(=O)Oc1ccccc1C(=O)O + O --> OC(=O)c1ccccc1O + CC(=O)O"
20796 -2.564 -1.886 -2.662 -4.718 -7.380 AB + CD --> AD + BC "CC(=O)Oc1ccccc1C(=O)O + O --> OC(=O)c1ccccc1O + CC(=O)O"
19675 -62.483 -62.709 -64.021 32.629 -31.392 AB + C --> AC + B "CC(=O)Oc1ccccc1C(=O)O xc{m06-2x} + [OH-] xc{m06-2x} --> CC(=O)Oc1ccccc1C(=O)[O-] xc{m06-2x} + O xc{m06-2x}"
17121 -59.866 -60.315 -61.722 38.458 -23.264 AB + C --> AC + B "CC(=O)Oc1ccccc1C(=O)O xc{pbe0} + [OH-] xc{pbe0} --> CC(=O)Oc1ccccc1C(=O)[O-] xc{pbe0} + O xc{pbe0}"
17113 -62.483 -62.710 -64.021 38.479 -25.543 AB + C --> AC + B "CC(=O)Oc1ccccc1C(=O)O xc{m06-2x} + [OH-] xc{m06-2x} --> CC(=O)Oc1ccccc1C(=O)[O-] xc{m06-2x} + O xc{m06-2x}"
17106 -57.969 -58.584 -58.981 41.419 -17.562 AB + C --> AC + B "CC(=O)Oc1ccccc1C(=O)O xc{pbe} + [OH-] xc{pbe} --> CC(=O)Oc1ccccc1C(=O)[O-] xc{pbe} + O xc{pbe}"
17103 -57.838 -58.382 -59.545 38.079 -21.466 AB + C --> AC + B "CC(=O)Oc1ccccc1C(=O)O xc{b3lyp} + [OH-] xc{b3lyp} --> CC(=O)Oc1ccccc1C(=O)[O-] xc{b3lyp} + O xc{b3lyp}"
14690 -53.074 -53.734 -55.811 0.000 -55.811 AB + C --> AC + B "CC(=O)Oc1ccccc1C(=O)O theory{pspw4} + [OH-] theory{pspw4} --> CC(=O)Oc1ccccc1C(=O)[O-] theory{pspw4} + O theory{pspw4}"
4292 -2.564 -1.918 -2.664 -4.938 -7.602 AB + CD --> AD + BC "CC(=O)Oc1ccccc1C(=O)O + O --> OC(=O)c1ccccc1O + CC(=O)O"
4291 -2.564 -1.918 -2.664 -4.938 -7.602 AB + CD --> AD + BC "CC(=O)Oc1ccccc1C(=O)O + O --> OC(=O)c1ccccc1O + CC(=O)O"
4290 -2.564 -1.918 -2.664 -4.938 -7.602 AB + CD --> AD + BC "CC(=O)Oc1ccccc1C(=O)O + O --> OC(=O)c1ccccc1O + CC(=O)O"
3118 23.956 27.044 38.752 -1.439 37.313 AB + CD --> CABD "Aspirin + Cl --> CC(=O)OC1=C(C=[CH]([CH2]=C1)Cl)C(=O)O"
1605 -35.473 -36.508 -37.938 42.370 4.432 AB + C --> AC + B "CC(=O)Oc1ccccc1C(=O)O + [OH-] --> [CH2-]C(=O)Oc1ccccc1C(=O)O + O"
1344 -45.511 -49.385 -61.575 45.023 -16.552 ABCD + E --> A + BC + DE "aspirin + hydroxide ^{-1} --> OC(=O)c1ccccc1[O] ^{-1} + [CH2][C]=O + O"
1266 -35.473 -36.508 -37.938 42.370 4.432 ABCD + E --> A + BC + DE "aspirin + hydroxide ^{-1} --> [CH2]C(=O)Oc1ccccc1C(=O)O ^{-1} + O"
1213 -52.728 -54.974 -57.479 52.164 -5.314 AB + C --> AC + B "CC(=O)Oc1ccccc1C(=O)O + [OH-] --> [CH2-]C(=O)Oc1ccccc1C(=O)O + O"
1212 -53.146 -53.750 -55.569 0.000 -55.569 AB + C --> AC + B "CC(=O)Oc1ccccc1C(=O)O theory{pspw4} + [OH-] theory{pspw4} --> CC(=O)Oc1ccccc1C(=O)[O-] theory{pspw4} + O theory{pspw4}"
813 23.956 27.044 38.752 -1.439 37.313 AB + CD --> CABD "Aspirin + Cl --> CC(=O)OC1=C(C=[CH]([CH2]=C1)Cl)C(=O)O"
804 -2.147 -3.176 -4.249 35.658 31.409 AB + C --> AC + B "Aspirin + hydroxide ^{-1} --> CC(=O)Oc1cc[c]cc1C(=O)O ^{-1} + O"
803 40.140 36.721 35.441 15.599 51.041 AB + C --> AC + B "Aspirin + [SH-] ^{-1} --> CC(=O)Oc1cc[c]cc1C(=O)O ^{-1} + S"
681 -2.564 -1.918 -2.664 -4.938 -7.602 AB + CD --> AD + BC "CC(=O)Oc1ccccc1C(=O)O + O --> OC(=O)c1ccccc1O + CC(=O)O"
All requests to Arrows were successful.
KEYWORDs -
reaction: :reaction
chinese_room: :chinese_room
molecule: :molecule
nmr: :nmr
predict: :predict
submitesmiles: :submitesmiles
nosubmitmissingesmiles
resubmitmissingesmiles
submitmachines: :submitmachines
useallentries
nomodelcorrect
eigenvalues: :eigenvalues
frequencies: :frequencies
nwoutput: :nwoutput
xyzfile: :xyzfile
alleigs: :alleigs
allfreqs: :allfreqs
reactionenumerate:
energytype:[erxn(gas) hrxn(gas) grxn(gas) delta_solvation grxn(aq)] :energytype
energytype:[kcal/mol kj/mol ev cm-1 ry hartree au] :energytype
tablereactions:
reaction: ... :reaction
reaction: ... :reaction
...
:tablereactions
tablemethods:
method: ... :method
method: ... :method
...
:tablemethods
:reactionenumerate
rotatebonds
xyzinput:
label: :label
xyzdata:
... xyz data ...
:xyzdata
:xyzinput
submitHf: :submitHf
nmrexp: :nmrexp
findreplace: old text | new text :findreplace
listnwjobs
fetchnwjob: :fetchnwjob
pushnwjob: :pushnwjob
printcsv: :printcsv
printeig: :printeig
printfreq: :printfreq
printxyz: :printxyz
printjobinfo: :printjobinfo
printnwout: :printnwout
badids: :badids
hup_string:
database:
table:
request_table:
listallesmiles
queuecheck
This software service and its documentation were developed at the Environmental Molecular Sciences Laboratory (EMSL) at Pacific Northwest National Laboratory, a multiprogram national laboratory, operated for the U.S. Department of Energy by Battelle under Contract Number DE-AC05-76RL01830. Support for this work was provided by the Department of Energy Office of Biological and Environmental Research, and Department of Defense environmental science and technology program (SERDP). THE SOFTWARE SERVICE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE SERVICE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE SERVICE.