 3D Periodic Editor 3D Molecular And Reaction Editor Expert Editor Microsoft Quantum Editor EMSL Aerosol Workshop Editor Manual
3D Periodic Editor 3D Molecular And Reaction Editor Expert Editor Microsoft Quantum Editor EMSL Aerosol Workshop Editor Manual Results from an EMSL Arrows Calculation
| EMSL Arrows is a revolutionary approach to materials and chemical simulations that uses NWChem and chemical computational databases to make materials and chemical modeling accessible via a broad spectrum of digital communications including posts to web APIs, social networks, and traditional email. |
Molecular modeling software has previously been extremely complex, making it prohibative to all but experts in the field, yet even experts can struggle to perform calculations. This service is designed to be used by experts and non-experts alike. Experts can carry out and keep track of large numbers of complex calculations with diverse levels of theories present in their workflows. Additionally, due to a streamlined and easy-to-use input, non-experts can carry out a wide variety of molecular modeling calculations previously not accessible to them.
The id(s) for emsiles = [CH]O theory{pspw4} xc{pbe} basis{100.0 Ry} solvation_type{None} ^{0} are: 76509
Use id=% instead of esmiles to print other entries.
mformula = C1H2O1
iupac = methanol
PubChem = 887
PubChem LCSS = 887
cas = 67-56-1
kegg = C00132 D02309
synonyms = methanol; methyl alcohol; 67-56-1; wood alcohol; carbinol; Wood spirit; Wood naphtha; Methylol; Methyl hydroxide; Pyroxylic spirit; Colonial Spirit; Columbian Spirit; Monohydroxymethane; Methylalkohol; Columbian spirits; Alcohol, methyl; Alcool methylique; MeOH; Methyl hydrate; CH3OH; Metanolo; Alcool metilico; Bieleski's solution; Colonial spirits; Metylowy alkohol; Pyroxylic spirits; Hydroxymethane; Freers Elm Arrester; Surflo-B17; Rcra waste number U154; Wilbur-Ellis Smut-Guard; Metanol [Spanish]; Metanolo [Italian]; Coat-B1400; Eureka Products, Criosine; Metanol; Caswell No. 552; Methylalkohol [German]; Spirit of wood; Alcool metilico [Italian]; Metylowy alkohol [Polish]; Alcool methylique [French]; X-Cide 402 Industrial Bactericide; HSDB 93; Ideal Concentrated Wood Preservative; Methyl alcohol [NF]; Eureka Products Criosine Disinfectant; NSC 85232; UN1230; CCRIS 2301; Alcohol,methyl; Pyro alcohol; AI3-00409; MetOH; RCRA waste no. U154; EPA Pesticide Chemical Code 053801; Methanol-water mixture; CHEBI:17790; NSC-85232; Y4S76JWI15; Methanol, anhydrous; MFCD00004595; Methyl alcohol (NF); NCGC00091172-01; Aqualine™ Solvent; Methanol-[17O]; DSSTox_CID_1731; Aqualine™ Titrant 5; DSSTox_RID_76297; DSSTox_GSID_21731; 170082-17-4; CH4O; Methanol, for HPLC, >=99.9%; Methanol, ACS reagent, >=99.8%; Methanol, or methyl alcohol [UN1230] [Flammable liquid, Poison]; CAS-67-56-1; MOH; EINECS 200-659-6; UNII-Y4S76JWI15; Methylalcohol; methly alcohol; Primary alcohol; methanol-; Wood; primary alcohols; Methanol cluster; Methanol NF; Nat. Methanol; a primary alcohol; Methanol LC-MS; Methanol, for HPLC; Methanol (Recovered); Methanol, ACS Grade; Solutions, Bieleski's; Methanol, HPLC grade; Methanol, LCMS grade; Columbian spirits; METHANOL [MI]; Hydroxymethylidyne radical; 3'-Hydroxystanozolol-D3; Methanol (Peptide Grade); Methanol, Histology Grade; bmse000294; Epitope ID:116865; METHANOL [USP-RS]; METHANOL [WHO-DD]; EC 200-659-6; Aqualine™ Solvent CM; Methanol Reagent Grade ACS; Methanol, or methyl alcohol; METHYL ALCOHOL [II]; Methanol, LR, >=99%; Methanol, SAJ special grade; Methanol, analytical standard; METHYL ALCOHOL [FCC]; WLN: Q1; Methanol HPLC Gradient Grade; Methanol, Environmental Grade; Aqualine™ Electrolyte A; CHEMBL14688; METHYL ALCOHOL [HSDB]; METHYL ALCOHOL [INCI]; ALCOHOL,METHYL [VANDF]; Methanol, anhydrous, 99.8%; Methanol, p.a., 99.8%; Methanol, p.a., 99.9%; Aqualine™ Electrolyte AG; Aqualine™ Electrolyte CG; METHANOL [EP MONOGRAPH]; METHYL ALCOHOL [MART.]; DTXSID2021731; Methanol, AR, >=99.5%; METHYL ALCOHOL [USP-RS]; CHEBI:15734; Methanol, NMR reference standard; Methanol, ultrapure, HPLC Grade; METHYL ALCOHOL (METHANOL); Methanol, 99.8%, ACS reagent; Methanol, anhydrous, >=99.5%; Methanol, low water for titration; Methanol GC, for residue analysis; Eriochrome™ Black T Solution; Methanol, Absolute - Acetone free; Methanol, low benzene, HPLC grade; Methanol, HPLC gradient, 99.9%; Methanol, or methyl alcohol [UN1230] [Flammable liquid, Poison]; NSC85232; Methanol, for HPLC, >=99.8%; Methanol, PRA grade, >=99.9%; Tox21_111094; Tox21_202523; Methanol, HPLC Plus, >=99.9%; AKOS000269045; Methanol, purification grade, 99.8%; UN 1230; Methanol, UHPLC, for mass spectrometry; ACETONE IMPURITY A [EP IMPURITY]; Methanol solution, technical grade, 95%; Methanol, >=99.8%, for chromatography; Methanol, SAJ first grade, >=99.5%; NCGC00260072-01; Methanol, JIS special grade, >=99.8%; Methanol, Laboratory Reagent, >=99.6%; Methanol, UV HPLC spectroscopic, 99.9%; Methanol, anhydrous, ZerO2(TM), 99.8%; Methanol, spectrophotometric grade, >=99%; FT-0623465; FT-0628297; FT-0628299; FT-0700908; FT-0700959; M0097; M0628; Methanol, ultrapure, Spectrophotometric Grade; C00132; D02309; Methanol, for HPLC, gradient grade, 99.93%; Methanol, suitable for determination of dioxins; Q14982; Methanol, for HPLC, gradient grade, >=99.9%; Methanol, glass distilled HRGC/HPLC trace grade; Methanol, low benzene, ACS reagent, = 99.8%; Methanol, low benzene, ACS reagent, >=99.8%; Methanol, ACS spectrophotometric grade, >=99.9%; Methanol HPLC, UV-IR min. 99.9% isocratic grade; Methanol, BioReagent, suitable for protein sequencing; Methanol, for HPLC, gradient grade, >=99.8% (GC); Methanol, HPLC Plus, >=99.9%, poly-coated bottles; Q27115113; Methanol solution, (Methanol:Acetonitrile 1:1 (v/v)); Methanol solution, contains 0.50 % (v/v) triethylamine; Methanol, Vetec(TM) reagent grade, anhydrous, >=99.8%; Methanol solution, (Methanol:Dichloromethane 1:1 (v/v)); Methanol, for residue analysis, suitable for 5000 per JIS; Moisture in methanol, 325 mg/kg, NIST(R) SRM(R) 8510; Moisture in methanol, 93 mg/kg, NIST(R) SRM(R) 8509; Methanol solution, (Methanol:Dimethyl sulfoxide 1:1 (v/v)); Methanol solution, contains 0.1 % (v/v) trifluoroacetic acid; Methanol solution, for protein sequence analysis, ~50% in H2O; Methanol with 0.1% trifluoroacetic acid, tested for UHPLC-MS; Methanol, >=99.8%, suitable for absorption spectrum analysis; Methanol, semiconductor grade PURANAL(TM) (Honeywell 17824); Methanol, p.a., ACS reagent, reag. ISO, reag. Ph. Eur., 99.9%; Methanol, puriss. p.a., absolute, ACS reagent, >=99.8% (GC); Methanol, semiconductor grade VLSI PURANAL(TM) (Honeywell 17744); Methanol, suitable for protein sequencing, BioReagent, >=99.93%; Methyl alcohol, United States Pharmacopeia (USP) Reference Standard; Methanol, Pharmaceutical Secondary Standard; Certified Reference Material; Methanol, puriss., meets analytical specification of Ph Eur, >=99.7% (GC); Methanol, suitable for 1000 per JIS, >=99.8%, for residue analysis; Methanol, suitable for 300 per JIS, >=99.8%, for residue analysis; (5beta,17beta)-17-Hydroxy-17-(methyl-d3)-2'H-androst-2-eno[3,2-c]pyrazol-5'(1'H)-one; Methanol solution, contains 0.1 % (v/v) trifluoroacetic acid, 5 % (v/v) water, for HPLC; Methanol solution, contains 0.10 % (v/v) trifluoroacetic acid, 10 % (v/v) water; Methanol solution, for HPLC, contains 10 % (v/v) water, 0.1 % (v/v) trifluoroacetic acid; Methanol, for HPLC, gradient grade, suitable as ACS-grade LC reagent, >=99.9%; Methanol, puriss. p.a., ACS reagent, reag. ISO, reag. Ph. Eur., >=99.8% (GC); Residual Solvent Class 2 - Methanol, United States Pharmacopeia (USP) Reference Standard; JandaJel(TM)-OH, 100-200 mesh, extent of labeling: 1.0 mmol/g OH loading, 2 % cross-linked; JandaJel(TM)-OH, 200-400 mesh, extent of labeling: 1.0 mmol/g OH loading, 2 % cross-linked; JandaJel(TM)-OH, 50-100 mesh, extent of labeling: 1.0 mmol/g OH loading, 2 % cross-linked; Methanol solution, contains 0.10 % (v/v) formic acid, UHPLC, for mass spectrometry, >=99.5%; Methanol solution, NMR reference standard, 4% in methanol-d4 (99.8 atom % D), NMR tube size 3 mm x 8 in.; Methanol solution, NMR reference standard, 4% in methanol-d4 (99.8 atom % D), NMR tube size 5 mm x 8 in.
Search Links to Other Online Resources (may not be available):
- Google Structure Search
- EPA CompTox Database
- Chemical Entities of Biological Interest (ChEBI)
- NIH ChemIDplus - A TOXNET DATABASE
- The Human Metabolome Database (HMDB)
- OECD eChemPortal
- Google Scholar
+==================================================+
|| Molecular Calculation ||
+==================================================+
Id = 76509
NWOutput = Link to NWChem Output (download)
Datafiles:
density.cube-980295-2023-1-22-1:37:1 (download)
lumo-restricted.cube-980295-2023-1-22-1:37:1 (download)
homo-restricted.cube-980295-2023-1-22-1:37:1 (download)
image_resset: api/image_reset/76509
Calculation performed by Eric Bylaska - bylaskamac
Numbers of cpus used for calculation = 8
Calculation walltime = 2283.700000 seconds (0 days 0 hours 38 minutes 3 seconds)
+----------------+
| Energetic Data |
+----------------+
Id = 76509
iupac = methanol
mformula = C1H2O1
inchi = InChI=1S/CH2O/c1-2/h1-2H
inchikey = UUOIGPDTLYBNGY-UHFFFAOYSA-N
esmiles = [CH]O theory{pspw4} xc{pbe} basis{100.0 Ry} solvation_type{None} ^{0}
calculation_type = ov
theory = pspw4
xc = pbe
basis = 100.0 Ry
charge,mult = 0 1
energy = -22.735801 Hartrees
enthalpy correct.= 0.029031 Hartrees
entropy = 53.740 cal/mol-K
solvation energy = 0.000 kcal/mol solvation_type = None
Sitkoff cavity dispersion = 1.613 kcal/mol
Honig cavity dispersion = 3.765 kcal/mol
ASA solvent accesible surface area = 150.613 Angstrom2
ASA solvent accesible volume = 144.138 Angstrom3
+-----------------+
| Structural Data |
+-----------------+
JSmol: an open-source HTML5 viewer for chemical structures in 3D
Number of Atoms = 4
Units are Angstrom for bonds and degrees for angles
Type I J K L M Value
----------- ----- ----- ----- ----- ----- ----------
1 Stretch C1 H2 1.11341
2 Stretch C1 O3 1.30399
3 Stretch O3 H4 0.96756
4 Bend H2 C1 O3 107.34635
5 Bend C1 O3 H4 115.63554
6 Dihedral H2 C1 O3 H4 0.15238
+-----------------+
| Eigenvalue Data |
+-----------------+
Id = 76509
iupac = methanol
mformula = C1H2O1
InChI = InChI=1S/CH2O/c1-2/h1-2H
smiles = [CH]O
esmiles = [CH]O theory{pspw4} xc{pbe} basis{100.0 Ry} solvation_type{None} ^{0}
theory = pspw4
xc = pbe
basis = 100.0 Ry
charge = 0
mult = 1
solvation_type = None
twirl webpage = TwirlMol Link
image webpage = GIF Image Link
Eigevalue Spectra
--- -- --- 0.95 eV
---- ----
----------
----------
---------- LUMO= -2.93 eV
HOMO= -5.10 eV ++++++++++
++++++++++
++++++++++
++++++++++
++++++++++
-28.19 eV ++++++++++
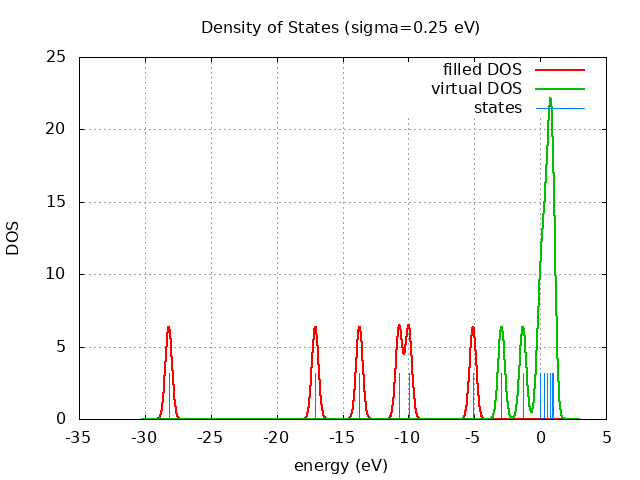
spin eig occ ---------------------------- restricted -5.10 2.00 restricted -9.98 2.00 restricted -10.71 2.00 restricted -13.72 2.00 restricted -17.07 2.00 restricted -28.19 2.00 restricted 0.95 0.00 restricted 0.90 0.00 restricted 0.77 0.00 restricted 0.55 0.00 restricted 0.27 0.00 restricted -0.03 0.00 restricted -1.31 0.00 restricted -2.93 0.00
+----------------------------------------+ | Vibrational Density of States Analysis | +----------------------------------------+ Total number of frequencies = 12 Total number of negative frequencies = 0 Number of lowest frequencies = 0 (frequency threshold = 500 ) Exact dos norm = 6.000 Generating vibrational DOS Generating model vibrational DOS to have a proper norm 10.00 6.00 0.00 6.00 50.00 6.00 0.00 6.00 100.00 6.00 0.00 6.00 Generating IR Spectra Writing vibrational density of states (DOS) to vdos.dat Writing model vibrational density of states (DOS_FIXED) to vdos-model.dat Writing IR spectra to irdos.dat Temperature= 298.15 zero-point correction to energy = 15.815 kcal/mol ( 0.025202) vibrational contribution to enthalpy correction = 15.848 kcal/mol ( 0.025256) vibrational contribution to Entropy = 0.133 cal/mol-k hindered rotor enthalpy correction = 0.000 kcal/mol ( 0.000000) hindered rotor entropy correction = 0.000 cal/mol-k


DOS sigma = 10.000000
- vibrational DOS enthalpy correction = 0.025256 kcal/mol ( 15.848 kcal/mol)
- model vibrational DOS enthalpy correction = 0.025256 kcal/mol ( 15.848 kcal/mol)
- vibrational DOS Entropy = 0.000000 ( 0.133 cal/mol-k)
- model vibrational DOS Entropy = 0.000000 ( 0.133 cal/mol-k)
- original gas Energy = -22.735801 (-14266.931 kcal/mol)
- original gas Enthalpy = -22.706770 (-14248.713 kcal/mol, delta= 0.000)
- unajusted DOS gas Enthalpy = -22.706770 (-14248.713 kcal/mol, delta= 0.000)
- model DOS gas Enthalpy = -22.706770 (-14248.713 kcal/mol, delta= 0.000)
- original gas Entropy = 0.000086 ( 53.740 cal/mol-k,delta= 0.000)
- unajusted DOS gas Entropy = 0.000086 ( 53.740 cal/mol-k,delta= 0.000)
- model DOS gas Entropy = 0.000086 ( 53.740 cal/mol-k,delta= 0.000)
- original gas Free Energy = -22.732304 (-14264.736 kcal/mol, delta= 0.000)
- unadjusted DOS gas Free Energy = -22.732304 (-14264.736 kcal/mol, delta= -0.000)
- model DOS gas Free Energy = -22.732304 (-14264.736 kcal/mol, delta= -0.000)
- original sol Free Energy = -22.732304 (-14264.736 kcal/mol)
- unadjusted DOS sol Free Energy = -22.732304 (-14264.736 kcal/mol)
- model DOS sol Free Energy = -22.732304 (-14264.736 kcal/mol)
DOS sigma = 50.000000
- vibrational DOS enthalpy correction = 0.025257 kcal/mol ( 15.849 kcal/mol)
- model vibrational DOS enthalpy correction = 0.025257 kcal/mol ( 15.849 kcal/mol)
- vibrational DOS Entropy = 0.000000 ( 0.136 cal/mol-k)
- model vibrational DOS Entropy = 0.000000 ( 0.136 cal/mol-k)
- original gas Energy = -22.735801 (-14266.931 kcal/mol)
- original gas Enthalpy = -22.706770 (-14248.713 kcal/mol, delta= 0.000)
- unajusted DOS gas Enthalpy = -22.706769 (-14248.713 kcal/mol, delta= 0.001)
- model DOS gas Enthalpy = -22.706769 (-14248.713 kcal/mol, delta= 0.001)
- original gas Entropy = 0.000086 ( 53.740 cal/mol-k,delta= 0.000)
- unajusted DOS gas Entropy = 0.000086 ( 53.743 cal/mol-k,delta= 0.003)
- model DOS gas Entropy = 0.000086 ( 53.743 cal/mol-k,delta= 0.003)
- original gas Free Energy = -22.732304 (-14264.736 kcal/mol, delta= 0.000)
- unadjusted DOS gas Free Energy = -22.732304 (-14264.736 kcal/mol, delta= -0.000)
- model DOS gas Free Energy = -22.732304 (-14264.736 kcal/mol, delta= -0.000)
- original sol Free Energy = -22.732304 (-14264.736 kcal/mol)
- unadjusted DOS sol Free Energy = -22.732304 (-14264.736 kcal/mol)
- model DOS sol Free Energy = -22.732304 (-14264.736 kcal/mol)
DOS sigma = 100.000000
- vibrational DOS enthalpy correction = 0.025260 kcal/mol ( 15.851 kcal/mol)
- model vibrational DOS enthalpy correction = 0.025260 kcal/mol ( 15.851 kcal/mol)
- vibrational DOS Entropy = 0.000000 ( 0.144 cal/mol-k)
- model vibrational DOS Entropy = 0.000000 ( 0.144 cal/mol-k)
- original gas Energy = -22.735801 (-14266.931 kcal/mol)
- original gas Enthalpy = -22.706770 (-14248.713 kcal/mol, delta= 0.000)
- unajusted DOS gas Enthalpy = -22.706766 (-14248.711 kcal/mol, delta= 0.003)
- model DOS gas Enthalpy = -22.706766 (-14248.711 kcal/mol, delta= 0.003)
- original gas Entropy = 0.000086 ( 53.740 cal/mol-k,delta= 0.000)
- unajusted DOS gas Entropy = 0.000086 ( 53.752 cal/mol-k,delta= 0.012)
- model DOS gas Entropy = 0.000086 ( 53.752 cal/mol-k,delta= 0.012)
- original gas Free Energy = -22.732304 (-14264.736 kcal/mol, delta= 0.000)
- unadjusted DOS gas Free Energy = -22.732305 (-14264.737 kcal/mol, delta= -0.001)
- model DOS gas Free Energy = -22.732305 (-14264.737 kcal/mol, delta= -0.001)
- original sol Free Energy = -22.732304 (-14264.736 kcal/mol)
- unadjusted DOS sol Free Energy = -22.732305 (-14264.737 kcal/mol)
- model DOS sol Free Energy = -22.732305 (-14264.737 kcal/mol)
Normal Mode Frequency (cm-1) IR Intensity (arbitrary)
----------- ---------------- ------------------------
1 0.000 1.174
2 0.000 1.607
3 0.000 4.651
4 0.000 3.546
5 0.000 11.206
6 0.000 8.622
7 1130.600 11.819
8 1228.390 11.417
9 1296.520 9.846
10 1470.140 16.716
11 2702.290 38.695
12 3239.770 0.701
No Hindered Rotor Data
+-------------------------------------+
| Reactions Contained in the Database |
+-------------------------------------+
Reactions Containing INCHIKEY = UUOIGPDTLYBNGY-UHFFFAOYSA-N
Reactionid Erxn(gas) Hrxn(gas) Grxn(gas) delta_Solv Grxn(aq) ReactionType Reaction
21449 63.528 61.951 52.757 -12.538 40.219 ACB --> AB + C "O[CH](=[Cl])=[Cl] ^{-1} mult{2} --> [CH]O + ClCl ^{-1} mult{2}"
20746 104.634 101.574 90.484 -4.258 86.226 ACB --> AB + C "C(Cl)(Cl)O xc{pbe0} --> [CH]O xc{pbe0} + ClCl xc{pbe0}"
19942 -45.003 -43.666 -33.977 24.838 -9.139 A + B --> AB "[CH]O xc{m06-2x} + [OH-] xc{m06-2x} --> O[CH-]O xc{m06-2x}"
19922 -63.673 -61.037 -59.306 8.933 -50.373 AB + C --> AC + B "[CH]Cl mult{3} xc{m06-2x} + [OH-] xc{m06-2x} --> [CH]O mult{3} xc{m06-2x} + [Cl-] xc{m06-2x}"
19311 95.717 92.684 81.648 -4.329 77.319 ACB --> AB + C "C(Cl)(Cl)O xc{b3lyp} --> [CH]O xc{b3lyp} + ClCl xc{b3lyp}"
17293 129.744 133.747 136.167 -138.305 -2.139 AB + C --> AC + B "[CH]=[Cl] ^{-1} mult{2} + [OH-] ^{-1} --> [CH]O + [Cl] ^{-2} mult{2}"
16813 93.406 90.381 79.349 -4.309 75.040 ACB --> AB + C "C(Cl)(Cl)O xc{b3lyp} --> [CH]O xc{b3lyp} + ClCl xc{b3lyp}"
15645 -54.690 -52.269 -50.531 14.094 -36.437 AB + C --> AC + B "[CH]Cl mult{3} theory{ccsd(t)} + [OH-] theory{ccsd(t)} --> [CH]O mult{3} theory{ccsd(t)} + [Cl-] theory{ccsd(t)}"
15620 51.116 50.660 43.626 -28.786 14.840 AB --> A + B "O[CH-]Cl theory{ccsd(t)} --> [CH]O mult{3} theory{ccsd(t)} + [Cl-] theory{ccsd(t)}"
15619 51.116 50.660 43.626 -28.786 14.840 AB --> A + B "O[CH-]Cl theory{ccsd(t)} --> [CH]O mult{3} theory{ccsd(t)} + [Cl-] theory{ccsd(t)}"
15611 13.491 11.518 3.583 -79.064 23.118 AB --> A + B "Cl[CH]O mult{3} theory{ccsd(t)} + SHE theory{ccsd(t)} --> [CH]O mult{3} theory{ccsd(t)} + [Cl-] theory{ccsd(t)}"
15610 13.491 11.518 3.583 -79.064 23.118 AB --> A + B "Cl[CH]O mult{3} theory{ccsd(t)} + SHE theory{ccsd(t)} --> [CH]O mult{3} theory{ccsd(t)} + [Cl-] theory{ccsd(t)}"
15576 -77.077 -81.749 -89.290 7.055 -82.235 CABD --> AB + CD "[CH]O mult{3} theory{ccsd(t)} --> [C-]#[O+] theory{ccsd(t)} + [HH] theory{ccsd(t)}"
15575 -77.077 -81.749 -89.290 7.055 -82.235 CABD --> AB + CD "[CH]O mult{3} theory{ccsd(t)} --> [C-]#[O+] theory{ccsd(t)} + [HH] theory{ccsd(t)}"
15427 50.345 49.875 42.673 -28.729 13.944 AB --> A + B "O[CH-]Cl xc{m06-2x} --> [CH]O mult{3} xc{m06-2x} + [Cl-] xc{m06-2x}"
15426 50.345 49.875 42.673 -28.729 13.944 AB --> A + B "O[CH-]Cl xc{m06-2x} --> [CH]O mult{3} xc{m06-2x} + [Cl-] xc{m06-2x}"
15425 25.921 24.966 17.805 -19.183 -1.378 AB --> A + B "O[CH-]Cl xc{pbe0} --> [CH]O mult{3} xc{pbe0} + [Cl-] xc{pbe0}"
15424 25.921 24.966 17.805 -19.183 -1.378 AB --> A + B "O[CH-]Cl xc{pbe0} --> [CH]O mult{3} xc{pbe0} + [Cl-] xc{pbe0}"
15423 31.476 30.772 23.549 -20.007 3.542 AB --> A + B "O[CH-]Cl xc{pbe} --> [CH]O mult{3} xc{pbe} + [Cl-] xc{pbe}"
15422 31.476 30.772 23.549 -20.007 3.542 AB --> A + B "O[CH-]Cl xc{pbe} --> [CH]O mult{3} xc{pbe} + [Cl-] xc{pbe}"
15421 48.508 48.051 41.017 -28.786 12.231 AB --> A + B "O[CH-]Cl xc{b3lyp} --> [CH]O mult{3} xc{b3lyp} + [Cl-] xc{b3lyp}"
15420 48.508 48.051 41.017 -28.786 12.231 AB --> A + B "O[CH-]Cl xc{b3lyp} --> [CH]O mult{3} xc{b3lyp} + [Cl-] xc{b3lyp}"
15419 28.457 27.619 20.060 0.000 20.060 AB --> A + B "O[CH-]Cl theory{pspw4} --> [CH]O mult{3} theory{pspw4} + [Cl-] theory{pspw4}"
15418 28.457 27.619 20.060 0.000 20.060 AB --> A + B "O[CH-]Cl theory{pspw4} --> [CH]O mult{3} theory{pspw4} + [Cl-] theory{pspw4}"
15398 -61.627 -68.926 -76.470 0.000 -76.470 CABD --> AB + CD "[CH]O mult{3} theory{pspw4} --> [C-]#[O+] theory{pspw4} + [HH] theory{pspw4}"
15397 -61.627 -68.926 -76.470 0.000 -76.470 CABD --> AB + CD "[CH]O mult{3} theory{pspw4} --> [C-]#[O+] theory{pspw4} + [HH] theory{pspw4}"
15333 -92.158 -87.095 -75.569 3.708 -71.861 AB + C --> ACB "[CH]O mult{3} xc{m06-2x} + O xc{m06-2x} --> C(O)O xc{m06-2x}"
15332 -92.158 -87.095 -75.569 3.708 -71.861 AB + C --> ACB "[CH]O mult{3} xc{m06-2x} + O xc{m06-2x} --> C(O)O xc{m06-2x}"
15331 -89.414 -84.347 -72.889 3.788 -69.102 AB + C --> ACB "[CH]O mult{3} xc{pbe0} + O xc{pbe0} --> C(O)O xc{pbe0}"
15330 -89.414 -84.347 -72.889 3.788 -69.102 AB + C --> ACB "[CH]O mult{3} xc{pbe0} + O xc{pbe0} --> C(O)O xc{pbe0}"
15329 -89.616 -84.731 -73.307 3.859 -69.448 AB + C --> ACB "[CH]O mult{3} xc{pbe} + O xc{pbe} --> C(O)O xc{pbe}"
15328 -89.616 -84.731 -73.307 3.859 -69.448 AB + C --> ACB "[CH]O mult{3} xc{pbe} + O xc{pbe} --> C(O)O xc{pbe}"
15327 -86.487 -81.139 -69.644 0.000 -69.644 AB + C --> ACB "[CH]O mult{3} theory{pspw4} + O theory{pspw4} --> C(O)O theory{pspw4}"
15326 -86.487 -81.139 -69.644 0.000 -69.644 AB + C --> ACB "[CH]O mult{3} theory{pspw4} + O theory{pspw4} --> C(O)O theory{pspw4}"
15213 -88.805 -83.657 -72.223 3.628 -68.595 AB + C --> ACB "[CH]O mult{3} xc{b3lyp} + O xc{b3lyp} --> C(O)O xc{b3lyp}"
15212 -88.805 -83.657 -72.223 3.628 -68.595 AB + C --> ACB "[CH]O mult{3} xc{b3lyp} + O xc{b3lyp} --> C(O)O xc{b3lyp}"
15201 -63.674 -61.037 -59.306 14.783 -44.524 AB + C --> AC + B "[CH]Cl mult{3} xc{m06-2x} + [OH-] xc{m06-2x} --> [CH]O mult{3} xc{m06-2x} + [Cl-] xc{m06-2x}"
15200 -60.946 -58.470 -56.737 14.094 -42.643 AB + C --> AC + B "[CH]Cl mult{3} xc{pbe0} + [OH-] xc{pbe0} --> [CH]O mult{3} xc{pbe0} + [Cl-] xc{pbe0}"
15199 -56.661 -54.446 -52.709 13.674 -39.036 AB + C --> AC + B "[CH]Cl mult{3} xc{pbe} + [OH-] xc{pbe} --> [CH]O mult{3} xc{pbe} + [Cl-] xc{pbe}"
15198 -59.899 -57.478 -55.740 14.094 -41.647 AB + C --> AC + B "[CH]Cl mult{3} xc{b3lyp} + [OH-] xc{b3lyp} --> [CH]O mult{3} xc{b3lyp} + [Cl-] xc{b3lyp}"
15197 -49.676 -47.251 -45.528 0.000 -45.528 AB + C --> AC + B "[CH]Cl mult{3} theory{pspw4} + [OH-] theory{pspw4} --> [CH]O mult{3} theory{pspw4} + [Cl-] theory{pspw4}"
15184 -70.395 -76.685 -84.222 6.996 -77.226 CABD --> AB + CD "[CH]O mult{3} xc{m06-2x} --> [C-]#[O+] xc{m06-2x} + [HH] xc{m06-2x}"
15183 -70.395 -76.685 -84.222 6.996 -77.226 CABD --> AB + CD "[CH]O mult{3} xc{m06-2x} --> [C-]#[O+] xc{m06-2x} + [HH] xc{m06-2x}"
15182 -60.503 -65.273 -72.833 6.956 -65.877 CABD --> AB + CD "[CH]O mult{3} xc{pbe0} --> [C-]#[O+] xc{pbe0} + [HH] xc{pbe0}"
15181 -60.503 -65.273 -72.833 6.956 -65.877 CABD --> AB + CD "[CH]O mult{3} xc{pbe0} --> [C-]#[O+] xc{pbe0} + [HH] xc{pbe0}"
15180 -63.715 -67.862 -75.408 6.976 -68.433 CABD --> AB + CD "[CH]O mult{3} xc{pbe} --> [C-]#[O+] xc{pbe} + [HH] xc{pbe}"
15179 -63.715 -67.862 -75.408 6.976 -68.433 CABD --> AB + CD "[CH]O mult{3} xc{pbe} --> [C-]#[O+] xc{pbe} + [HH] xc{pbe}"
15176 -68.616 -73.269 -80.810 7.055 -73.755 CABD --> AB + CD "[CH]O mult{3} --> [C-]#[O+] + [HH]"
15175 -68.616 -73.269 -80.810 7.055 -73.755 CABD --> AB + CD "[CH]O mult{3} --> [C-]#[O+] + [HH]"
15111 -56.637 -53.039 -44.594 0.000 -44.594 A + B --> AB "[CH]O theory{pspw4} xc{pbe0} + [OH-] theory{pspw4} xc{pbe0} --> O[CH-]O theory{pspw4} xc{pbe0}"
15104 -48.211 -59.794 -67.495 0.000 -67.495 CABD --> AB + CD "[CH]O theory{pspw4} xc{pbe0} --> [C][O] theory{pspw4} xc{pbe0} + [HH] theory{pspw4} xc{pbe0}"
15103 -48.211 -59.794 -67.495 0.000 -67.495 CABD --> AB + CD "[CH]O theory{pspw4} xc{pbe0} --> [C][O] theory{pspw4} xc{pbe0} + [HH] theory{pspw4} xc{pbe0}"
15099 11.952 9.903 1.710 0.000 100.310 AB --> A + B "Cl[CH]O theory{pspw4} xc{pbe0} + SHE theory{pspw4} xc{pbe0} --> [CH]O mult{3} theory{pspw4} xc{pbe0} + [Cl-] theory{pspw4} xc{pbe0}"
15098 11.952 9.903 1.710 0.000 100.310 AB --> A + B "Cl[CH]O theory{pspw4} xc{pbe0} + SHE theory{pspw4} xc{pbe0} --> [CH]O mult{3} theory{pspw4} xc{pbe0} + [Cl-] theory{pspw4} xc{pbe0}"
15096 0.643 -0.668 -8.713 0.000 89.887 AB --> A + B "Cl[CH]O theory{pspw4} xc{pbe0} + SHE theory{pspw4} xc{pbe0} --> [CH]O theory{pspw4} xc{pbe0} + [Cl-] theory{pspw4} xc{pbe0}"
15095 0.643 -0.668 -8.713 0.000 89.887 AB --> A + B "Cl[CH]O theory{pspw4} xc{pbe0} + SHE theory{pspw4} xc{pbe0} --> [CH]O theory{pspw4} xc{pbe0} + [Cl-] theory{pspw4} xc{pbe0}"
15060 97.391 94.807 83.645 -1.096 82.549 ACB --> AB + C "C(Cl)(Cl)O xc{m06-2x} --> [CH]O xc{m06-2x} + ClCl xc{m06-2x}"
15059 104.634 101.576 90.485 -4.328 86.158 ACB --> AB + C "C(Cl)(Cl)O xc{pbe0} --> [CH]O xc{pbe0} + ClCl xc{pbe0}"
15058 102.064 98.971 88.002 -4.199 83.804 ACB --> AB + C "C(Cl)(Cl)O xc{pbe} --> [CH]O xc{pbe} + ClCl xc{pbe}"
15057 95.717 92.686 81.648 -4.230 77.418 ACB --> AB + C "C(Cl)(Cl)O xc{b3lyp} --> [CH]O xc{b3lyp} + ClCl xc{b3lyp}"
14908 -58.586 -54.884 -45.332 0.000 -45.332 A + B --> AB "[CH]O theory{pspw4} + [OH-] theory{pspw4} --> O[CH-]O theory{pspw4}"
14899 -48.323 -56.201 -63.898 0.000 -63.898 CABD --> AB + CD "[CH]O theory{pspw4} --> [C][O] theory{pspw4} + [HH] theory{pspw4}"
14898 -48.323 -56.201 -63.898 0.000 -63.898 CABD --> AB + CD "[CH]O theory{pspw4} --> [C][O] theory{pspw4} + [HH] theory{pspw4}"
14897 -46.236 -53.573 -61.308 8.389 -52.919 CABD --> AB + CD "[CH]O xc{m06-2x} --> [C][O] xc{m06-2x} + [HH] xc{m06-2x}"
14896 -46.236 -53.573 -61.308 8.389 -52.919 CABD --> AB + CD "[CH]O xc{m06-2x} --> [C][O] xc{m06-2x} + [HH] xc{m06-2x}"
14895 -45.897 -51.322 -59.047 11.511 -47.536 CABD --> AB + CD "[CH]O xc{pbe0} --> [C][O] xc{pbe0} + [HH] xc{pbe0}"
14894 -45.897 -51.322 -59.047 11.511 -47.536 CABD --> AB + CD "[CH]O xc{pbe0} --> [C][O] xc{pbe0} + [HH] xc{pbe0}"
14893 -46.502 -51.156 -58.874 11.229 -47.644 CABD --> AB + CD "[CH]O xc{pbe} --> [C][O] xc{pbe} + [HH] xc{pbe}"
14892 -46.502 -51.156 -58.874 11.229 -47.644 CABD --> AB + CD "[CH]O xc{pbe} --> [C][O] xc{pbe} + [HH] xc{pbe}"
14891 -45.003 -43.666 -33.977 30.688 -3.289 A + B --> AB "[CH]O xc{m06-2x} + [OH-] xc{m06-2x} --> O[CH-]O xc{m06-2x}"
14890 -50.652 -49.258 -39.587 35.469 -4.119 A + B --> AB "[CH]O xc{pbe0} + [OH-] xc{pbe0} --> O[CH-]O xc{pbe0}"
14889 -52.411 -51.426 -41.795 40.280 -1.516 A + B --> AB "[CH]O xc{pbe} + [OH-] xc{pbe} --> O[CH-]O xc{pbe}"
14888 -44.933 -43.612 -33.985 36.071 2.087 A + B --> AB "[CH]O xc{b3lyp} + [OH-] xc{b3lyp} --> O[CH-]O xc{b3lyp}"
14871 12.832 10.978 3.025 0.000 101.625 AB --> A + B "Cl[CH]O theory{pspw4} + SHE theory{pspw4} --> [CH]O mult{3} theory{pspw4} + [Cl-] theory{pspw4}"
14870 12.832 10.978 3.025 0.000 101.625 AB --> A + B "Cl[CH]O theory{pspw4} + SHE theory{pspw4} --> [CH]O mult{3} theory{pspw4} + [Cl-] theory{pspw4}"
14869 7.713 5.793 -2.294 -79.434 16.872 AB --> A + B "Cl[CH]O xc{m06-2x} + SHE xc{m06-2x} --> [CH]O mult{3} xc{m06-2x} + [Cl-] xc{m06-2x}"
14868 7.713 5.793 -2.294 -79.434 16.872 AB --> A + B "Cl[CH]O xc{m06-2x} + SHE xc{m06-2x} --> [CH]O mult{3} xc{m06-2x} + [Cl-] xc{m06-2x}"
14867 10.191 8.224 0.260 -79.572 19.288 AB --> A + B "Cl[CH]O xc{pbe0} + SHE xc{pbe0} --> [CH]O mult{3} xc{pbe0} + [Cl-] xc{pbe0}"
14866 10.191 8.224 0.260 -79.572 19.288 AB --> A + B "Cl[CH]O xc{pbe0} + SHE xc{pbe0} --> [CH]O mult{3} xc{pbe0} + [Cl-] xc{pbe0}"
14865 11.705 9.776 1.954 -78.885 21.669 AB --> A + B "Cl[CH]O xc{pbe} + SHE xc{pbe} --> [CH]O mult{3} xc{pbe} + [Cl-] xc{pbe}"
14864 11.705 9.776 1.954 -78.885 21.669 AB --> A + B "Cl[CH]O xc{pbe} + SHE xc{pbe} --> [CH]O mult{3} xc{pbe} + [Cl-] xc{pbe}"
14863 3.901 1.928 -6.007 -79.064 13.529 AB --> A + B "Cl[CH]O xc{b3lyp} + SHE xc{b3lyp} --> [CH]O mult{3} xc{b3lyp} + [Cl-] xc{b3lyp}"
14862 3.901 1.928 -6.007 -79.064 13.529 AB --> A + B "Cl[CH]O xc{b3lyp} + SHE xc{b3lyp} --> [CH]O mult{3} xc{b3lyp} + [Cl-] xc{b3lyp}"
14847 -0.472 -1.747 -9.548 0.000 89.052 AB --> A + B "Cl[CH]O theory{pspw4} + SHE theory{pspw4} --> [CH]O theory{pspw4} + [Cl-] theory{pspw4}"
14846 -0.472 -1.747 -9.548 0.000 89.052 AB --> A + B "Cl[CH]O theory{pspw4} + SHE theory{pspw4} --> [CH]O theory{pspw4} + [Cl-] theory{pspw4}"
14845 -16.445 -17.319 -25.208 -80.828 -7.436 AB --> A + B "Cl[CH]O xc{m06-2x} + SHE xc{m06-2x} --> [CH]O xc{m06-2x} + [Cl-] xc{m06-2x}"
14844 -16.445 -17.319 -25.208 -80.828 -7.436 AB --> A + B "Cl[CH]O xc{m06-2x} + SHE xc{m06-2x} --> [CH]O xc{m06-2x} + [Cl-] xc{m06-2x}"
14843 -4.415 -5.727 -13.527 -84.127 0.946 AB --> A + B "Cl[CH]O xc{pbe0} + SHE xc{pbe0} --> [CH]O xc{pbe0} + [Cl-] xc{pbe0}"
14842 -4.415 -5.727 -13.527 -84.127 0.946 AB --> A + B "Cl[CH]O xc{pbe0} + SHE xc{pbe0} --> [CH]O xc{pbe0} + [Cl-] xc{pbe0}"
14841 -5.509 -6.930 -14.581 -83.138 0.881 AB --> A + B "Cl[CH]O xc{pbe} + SHE xc{pbe} --> [CH]O xc{pbe} + [Cl-] xc{pbe}"
14840 -5.509 -6.930 -14.581 -83.138 0.881 AB --> A + B "Cl[CH]O xc{pbe} + SHE xc{pbe} --> [CH]O xc{pbe} + [Cl-] xc{pbe}"
14810 -49.894 -55.246 -62.961 11.400 -51.562 CABD --> AB + CD "[CH]O xc{b3lyp} --> [C][O] xc{b3lyp} + [HH] xc{b3lyp}"
14809 -49.894 -55.246 -62.961 11.400 -51.562 CABD --> AB + CD "[CH]O xc{b3lyp} --> [C][O] xc{b3lyp} + [HH] xc{b3lyp}"
14808 -14.821 -16.094 -23.856 -83.409 -8.665 AB --> A + B "Cl[CH]O xc{b3lyp} + SHE xc{b3lyp} --> [CH]O xc{b3lyp} + [Cl-] xc{b3lyp}"
14807 -14.821 -16.094 -23.856 -83.409 -8.665 AB --> A + B "Cl[CH]O xc{b3lyp} + SHE xc{b3lyp} --> [CH]O xc{b3lyp} + [Cl-] xc{b3lyp}"
All requests to Arrows were successful.
KEYWORDs -
reaction: :reaction
chinese_room: :chinese_room
molecule: :molecule
nmr: :nmr
predict: :predict
submitesmiles: :submitesmiles
nosubmitmissingesmiles
resubmitmissingesmiles
submitmachines: :submitmachines
useallentries
nomodelcorrect
eigenvalues: :eigenvalues
frequencies: :frequencies
nwoutput: :nwoutput
xyzfile: :xyzfile
alleigs: :alleigs
allfreqs: :allfreqs
reactionenumerate:
energytype:[erxn(gas) hrxn(gas) grxn(gas) delta_solvation grxn(aq)] :energytype
energytype:[kcal/mol kj/mol ev cm-1 ry hartree au] :energytype
tablereactions:
reaction: ... :reaction
reaction: ... :reaction
...
:tablereactions
tablemethods:
method: ... :method
method: ... :method
...
:tablemethods
:reactionenumerate
rotatebonds
xyzinput:
label: :label
xyzdata:
... xyz data ...
:xyzdata
:xyzinput
submitHf: :submitHf
nmrexp: :nmrexp
findreplace: old text | new text :findreplace
listnwjobs
fetchnwjob: :fetchnwjob
pushnwjob: :pushnwjob
printcsv: :printcsv
printeig: :printeig
printfreq: :printfreq
printxyz: :printxyz
printjobinfo: :printjobinfo
printnwout: :printnwout
badids: :badids
hup_string:
database:
table:
request_table:
listallesmiles
queuecheck
This software service and its documentation were developed at the Environmental Molecular Sciences Laboratory (EMSL) at Pacific Northwest National Laboratory, a multiprogram national laboratory, operated for the U.S. Department of Energy by Battelle under Contract Number DE-AC05-76RL01830. Support for this work was provided by the Department of Energy Office of Biological and Environmental Research, and Department of Defense environmental science and technology program (SERDP). THE SOFTWARE SERVICE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE SERVICE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE SERVICE.